Benign prostatic hyperplasia (BPH) treatments have changed little over many years and do not directly address the underlying cause. Because BPH is characterised by uncontrolled cell growth, the chromosomal telomeres should be eroded in the reported absence or low levels of telomerase activity, but this is not observed. We investigated the telomere biology of cell subpopulations from BPH patients undergoing transurethral resection of prostate (TURP). Measurement of TERC, TERT, and telomerase activity revealed that only the epithelial stem-like and progenitor fractions expressed high levels of telomerase activity (p < 0.01) and individual enzyme components (p < 0.01). Telomerase activity and TERT expression were not detected in stromal cells. Telomere length measurements reflected this activity, although the average telomere length of (telomerase-negative) luminal cells was equivalent to that of telomerase-expressing stem/progenitor cells. Immunohistochemical analysis of patient-derived BPH arrays identified distinct areas of luminal hyperproliferation, basal hyperproliferation, and basal–luminal hyperproliferation, suggesting that basal and luminal cells can proliferate independently of each other. We propose a separate lineage for the luminal and basal cell components in BPH.
Patient summary
We unexpectedly found an enzyme called telomerase in the cells that maintain benign prostatic hyperplasia (BPH), suggesting that telomerase inhibitors could be used to alleviate BPH symptoms.
Morbidity and loss of quality of life due to benign prostatic hyperplasia (BPH) are relatively high. There are >210 million cases of BPH worldwide [1], and the histologic prevalence of BPH is approximately 80% at the age of 80 yr. BPH is characterised by uncontrolled but noninvasive growth of a variety of cell types in the prostate, ranging in frequency from adenomyofibromatous, fibromuscular, fibroadenomatous, stromal, to rare muscular compositions. Medical treatments for BPH include targeting of the smooth muscle cells by α1-adrenergic receptor inhibitor and use of steroid 5α-reductase inhibitors to target the androgen-signalling axis. These treatments take several months to show urinary flow rate benefits but perhaps are ineffective in targeting androgen-independent basal cell proliferation. Surgical intervention by transurethral resection relieves urinary flow obstruction.
In normal somatic cells, repeated cell divisions result in an erosion of the repetitive terminal DNA sequences, the telomeres. The ultimate result of extreme telomere shortening is stress signal–mediated apoptosis or senescence. An understanding of telomerase biology is crucial to comprehend the proliferative dynamics of a tissue. Increased telomerase can be detected in approximately 70–90% of human cancers, including prostate cancer [2]. Although BPH tissue also exhibits a proliferative index two to three times higher than normal prostate [3], telomerase remains undetectable or is only sporadically expressed [2]. The inability to detect telomerase in non–stem-like cells, including the differentiated basal and luminal cells (>95% of the lesion), could have been interpreted as absent or sporadic telomerase expression or activity patterns in previous whole-biopsy analyses [2].
We assessed expression of the RNA (TERC) and protein (TERT) components of telomerase enzyme, telomerase activity, and telomere length in uncultured human BPH-derived stem-like cells (SCs: Lin−/CD31−/EpCAM+/CD133+/CD44+/α2β1hi), transit amplifying cells (TAs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1hi), committed basal cells (CBs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1lo), luminal cells (LCs: Lin−/CD31−/EpCAM+/CD44−/CD24+), and stromal cells (Lin−/CD31−/EpCAM−) (Supplementary Fig. 1, Supplement 1) [4] and [5] freshly purified from BPH tissues (Supplementary Table 1). The expression of NANOG was also higher in SCs (Supplementary Fig. 2). Previously, we reported that TERT expression was undetectable in normal prostate epithelial SCs and TAs [6]. In human BPH, however, the SCs and TAs exhibited significant overexpression of both TERC and TERT compared with the more differentiated CBs and LCs and stromal cells (Fig. 1a and 1b). Telomerase activity was also exclusively restricted to the SC/TA compartment, apart from one CB sample (Fig. 1c). All biopsies were specifically taken from the transitional zone with no detectable prostate cancer lesions on histology from patients with serum prostate-specific antigen levels <4 ng/ml to reduce the possibility of contamination with (telomerase-positive) cancer cells (Supplementary Table 1). TERT expression and telomerase activity were undetectable in stromal cells, indicating that telomerase is not essential for stromal hyperproliferation in BPH—another intriguing disparity in earlier data.
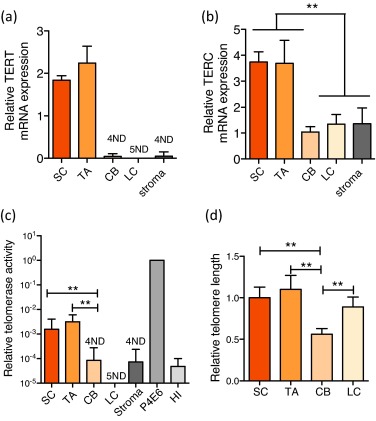
Epithelial cell telomere length measurements revealed that telomerase-expressing SCs and TAs had longer telomeres than those of their immediate differentiated progeny, the telomerase-negative CBs (p < 0.01) (Fig. 1d). The relative telomere length of LCs, however, was almost equivalent to that of SCs and TAs and was always longer than telomere lengths of patient-matched CBs (p < 0.01). Because LCs do not exhibit any detectable telomerase activity (Fig. 1c) and are believed to be derived from basal cells, the paradoxical observation of longer LC telomeres compared with CBs suggests that basal and luminal cells may be derived from distinct SC progenitors.
We stained BPH sections to determine basal cell content (using TP63) and luminal cell content (using NKX3.1) (Fig. 2a) and showed that basal and luminal hyperproliferation can occur together and even independently of each other because the basal:luminal ratio was highly variable, ranging from 0.22 to 2.08 (Fig. 2b, Supplementary Fig. 3), providing further evidence that basal and luminal cells may indeed be derived from different progenitors.
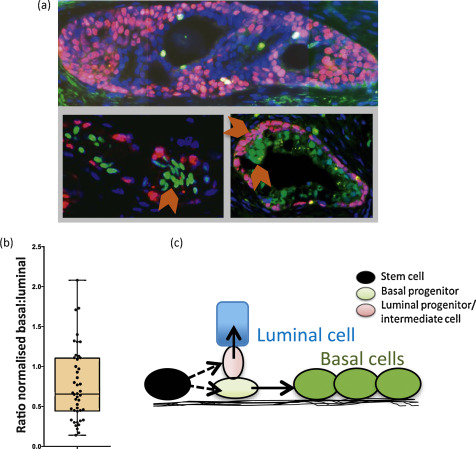
In human prostate, we have previously shown that SCs differentiate into CB cells via TA cells [4], unlike mouse cells, in which the situation is less clear [7]. The lineage of LCs, however, has not been determined in humans, although in vitro we can readily derive luminal cells from basal cells under specific culture conditions [5]. We now propose that basal and luminal cell lineages may develop separately in BPH and speculate that the prostate epithelial SCs can asymmetrically differentiate [8] into distinct basal and luminal progenitor cells, as shown in Figure 2c. In many other stem cell systems, the spindle orientation of progenitor progenies can determine their ultimate identity. The two-progenitor model may assist in understanding BPH pathogenesis, in which each progenitor and its progeny can expand independently or simultaneously to give rise to well-documented [9] areas of basal hyperproliferation, luminal hyperproliferation, or proliferation of both compartments, as shown in Figure 2a. If a basal progenitor cell differentiates into CBs, which retain limited proliferative potential in the absence of detectable telomerase activity, this will result in telomere shortening in CBs, for example, in basal cell hyperplasia, as shown in Figure 2a. In contrast, a luminal progenitor cell population expressing telomerase could expand and give rise to terminally differentiated luminal cells, which do not replicate, maintaining a telomere length similar to that of the luminal progenitors. Our data is in agreement with both Bonkhoff et al [10] and McNeal et al [11] with respect to the existence of a proliferative basal compartment in which the potential luminal progenitors derived from basal SCs exhibit characteristics of both basal and luminal phenotypes, analogous to castration-resistant Nkx3.1-expressing cells in mice [12]. The second implication of our model is that local inhibition of telomerase in BPH could be an alternative therapeutic strategy, as it will inhibit the proliferation both luminal and basal epithelial progenitors but perhaps not that of stromal cells. Because it is suggested that these progenitors function as a reservoir of proliferating epithelial cells in BPH, utilisation of novel and specific telomerase inhibitors such as Imetelstat (Geron Corp, Menlo Park, CA, USA) could prove to be a good adjuvant therapy for the management of BPH.
Author contributions: Norman J. Maitland had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Study concept and design: Maitland, Rane.
Acquisition of data: Rane, Frame, Greener.
Analysis and interpretation of data: Maitland, Rane, Frame.
Drafting of the manuscript: Rane.
Critical revision of the manuscript for important intellectual content: Maitland, Rane, Collins, Berney, Frame.
Statistical analysis: Rane, Frame.
Obtaining funding: Maitland.
Administrative, technical, or material support: Mann, Simms, Collins, Berney.
Supervision: Maitland.
Other (specify): None.
Financial disclosures: Norman J. Maitland certifies that all conflicts of interest, including specific financial interests and relationships and affiliations relevant to the subject matter or materials discussed in the manuscript (eg, employment/affiliation, grants or funding, consultancies, honoraria, stock ownership or options, expert testimony, royalties, or patents filed, received, or pending), are the following: None.
Funding/Support and role of the sponsor: The work was funded by Prostate Cancer UK (G2012-37) and PRO-NEST Marie-Curie network (J.K.R.), and Yorkshire Cancer Research (grant Y257PA, F.M.F. and N.J.M.). The human prostate tissue bank was funded by the Orchid Trust (D.B.). The sponsors were involved in the design and conduct of the study and collection and analysis of the data.
Acknowledgements: We would like to thank all of the patients and the urology surgeons L. Coombes, G. Cooksey and J. Hetherington (Castle Hill Hospital, Cottingham, UK). Thank you also to Megan Nash for immunofluorescence staining and counting and to Jerry Shay (University of Texas Southwestern, Dallas, TX, USA) for providing invaluable guidance on the telomerase and telomere measurement experiments. Davide Pellacani provided many useful discussions concerning the work.
Morbidity and loss of quality of life due to benign prostatic hyperplasia (BPH) are relatively high. There are >210 million cases of BPH worldwide [1], and the histologic prevalence of BPH is approximately 80% at the age of 80 yr. BPH is characterised by uncontrolled but noninvasive growth of a variety of cell types in the prostate, ranging in frequency from adenomyofibromatous, fibromuscular, fibroadenomatous, stromal, to rare muscular compositions. Medical treatments for BPH include targeting of the smooth muscle cells by α1-adrenergic receptor inhibitor and use of steroid 5α-reductase inhibitors to target the androgen-signalling axis. These treatments take several months to show urinary flow rate benefits but perhaps are ineffective in targeting androgen-independent basal cell proliferation. Surgical intervention by transurethral resection relieves urinary flow obstruction.
In normal somatic cells, repeated cell divisions result in an erosion of the repetitive terminal DNA sequences, the telomeres. The ultimate result of extreme telomere shortening is stress signal–mediated apoptosis or senescence. An understanding of telomerase biology is crucial to comprehend the proliferative dynamics of a tissue. Increased telomerase can be detected in approximately 70–90% of human cancers, including prostate cancer [2]. Although BPH tissue also exhibits a proliferative index two to three times higher than normal prostate [3], telomerase remains undetectable or is only sporadically expressed [2]. The inability to detect telomerase in non–stem-like cells, including the differentiated basal and luminal cells (>95% of the lesion), could have been interpreted as absent or sporadic telomerase expression or activity patterns in previous whole-biopsy analyses [2].
We assessed expression of the RNA (TERC) and protein (TERT) components of telomerase enzyme, telomerase activity, and telomere length in uncultured human BPH-derived stem-like cells (SCs: Lin−/CD31−/EpCAM+/CD133+/CD44+/α2β1hi), transit amplifying cells (TAs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1hi), committed basal cells (CBs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1lo), luminal cells (LCs: Lin−/CD31−/EpCAM+/CD44−/CD24+), and stromal cells (Lin−/CD31−/EpCAM−) (Supplementary Fig. 1, Supplement 1) [4] and [5] freshly purified from BPH tissues (Supplementary Table 1). The expression of NANOG was also higher in SCs (Supplementary Fig. 2). Previously, we reported that TERT expression was undetectable in normal prostate epithelial SCs and TAs [6]. In human BPH, however, the SCs and TAs exhibited significant overexpression of both TERC and TERT compared with the more differentiated CBs and LCs and stromal cells (Fig. 1a and 1b). Telomerase activity was also exclusively restricted to the SC/TA compartment, apart from one CB sample (Fig. 1c). All biopsies were specifically taken from the transitional zone with no detectable prostate cancer lesions on histology from patients with serum prostate-specific antigen levels <4 ng/ml to reduce the possibility of contamination with (telomerase-positive) cancer cells (Supplementary Table 1). TERT expression and telomerase activity were undetectable in stromal cells, indicating that telomerase is not essential for stromal hyperproliferation in BPH—another intriguing disparity in earlier data.

Epithelial cell telomere length measurements revealed that telomerase-expressing SCs and TAs had longer telomeres than those of their immediate differentiated progeny, the telomerase-negative CBs (p < 0.01) (Fig. 1d). The relative telomere length of LCs, however, was almost equivalent to that of SCs and TAs and was always longer than telomere lengths of patient-matched CBs (p < 0.01). Because LCs do not exhibit any detectable telomerase activity (Fig. 1c) and are believed to be derived from basal cells, the paradoxical observation of longer LC telomeres compared with CBs suggests that basal and luminal cells may be derived from distinct SC progenitors.
We stained BPH sections to determine basal cell content (using TP63) and luminal cell content (using NKX3.1) (Fig. 2a) and showed that basal and luminal hyperproliferation can occur together and even independently of each other because the basal:luminal ratio was highly variable, ranging from 0.22 to 2.08 (Fig. 2b, Supplementary Fig. 3), providing further evidence that basal and luminal cells may indeed be derived from different progenitors.

In human prostate, we have previously shown that SCs differentiate into CB cells via TA cells [4], unlike mouse cells, in which the situation is less clear [7]. The lineage of LCs, however, has not been determined in humans, although in vitro we can readily derive luminal cells from basal cells under specific culture conditions [5]. We now propose that basal and luminal cell lineages may develop separately in BPH and speculate that the prostate epithelial SCs can asymmetrically differentiate [8] into distinct basal and luminal progenitor cells, as shown in Figure 2c. In many other stem cell systems, the spindle orientation of progenitor progenies can determine their ultimate identity. The two-progenitor model may assist in understanding BPH pathogenesis, in which each progenitor and its progeny can expand independently or simultaneously to give rise to well-documented [9] areas of basal hyperproliferation, luminal hyperproliferation, or proliferation of both compartments, as shown in Figure 2a. If a basal progenitor cell differentiates into CBs, which retain limited proliferative potential in the absence of detectable telomerase activity, this will result in telomere shortening in CBs, for example, in basal cell hyperplasia, as shown in Figure 2a. In contrast, a luminal progenitor cell population expressing telomerase could expand and give rise to terminally differentiated luminal cells, which do not replicate, maintaining a telomere length similar to that of the luminal progenitors. Our data is in agreement with both Bonkhoff et al [10] and McNeal et al [11] with respect to the existence of a proliferative basal compartment in which the potential luminal progenitors derived from basal SCs exhibit characteristics of both basal and luminal phenotypes, analogous to castration-resistant Nkx3.1-expressing cells in mice [12]. The second implication of our model is that local inhibition of telomerase in BPH could be an alternative therapeutic strategy, as it will inhibit the proliferation both luminal and basal epithelial progenitors but perhaps not that of stromal cells. Because it is suggested that these progenitors function as a reservoir of proliferating epithelial cells in BPH, utilisation of novel and specific telomerase inhibitors such as Imetelstat (Geron Corp, Menlo Park, CA, USA) could prove to be a good adjuvant therapy for the management of BPH.
Author contributions: Norman J. Maitland had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Study concept and design: Maitland, Rane.
Acquisition of data: Rane, Frame, Greener.
Analysis and interpretation of data: Maitland, Rane, Frame.
Drafting of the manuscript: Rane.
Critical revision of the manuscript for important intellectual content: Maitland, Rane, Collins, Berney, Frame.
Statistical analysis: Rane, Frame.
Obtaining funding: Maitland.
Administrative, technical, or material support: Mann, Simms, Collins, Berney.
Supervision: Maitland.
Other (specify): None.
Financial disclosures: Norman J. Maitland certifies that all conflicts of interest, including specific financial interests and relationships and affiliations relevant to the subject matter or materials discussed in the manuscript (eg, employment/affiliation, grants or funding, consultancies, honoraria, stock ownership or options, expert testimony, royalties, or patents filed, received, or pending), are the following: None.
Funding/Support and role of the sponsor: The work was funded by Prostate Cancer UK (G2012-37) and PRO-NEST Marie-Curie network (J.K.R.), and Yorkshire Cancer Research (grant Y257PA, F.M.F. and N.J.M.). The human prostate tissue bank was funded by the Orchid Trust (D.B.). The sponsors were involved in the design and conduct of the study and collection and analysis of the data.
Acknowledgements: We would like to thank all of the patients and the urology surgeons L. Coombes, G. Cooksey and J. Hetherington (Castle Hill Hospital, Cottingham, UK). Thank you also to Megan Nash for immunofluorescence staining and counting and to Jerry Shay (University of Texas Southwestern, Dallas, TX, USA) for providing invaluable guidance on the telomerase and telomere measurement experiments. Davide Pellacani provided many useful discussions concerning the work.
Morbidity and loss of quality of life due to benign prostatic hyperplasia (BPH) are relatively high. There are >210 million cases of BPH worldwide [1], and the histologic prevalence of BPH is approximately 80% at the age of 80 yr. BPH is characterised by uncontrolled but noninvasive growth of a variety of cell types in the prostate, ranging in frequency from adenomyofibromatous, fibromuscular, fibroadenomatous, stromal, to rare muscular compositions. Medical treatments for BPH include targeting of the smooth muscle cells by α1-adrenergic receptor inhibitor and use of steroid 5α-reductase inhibitors to target the androgen-signalling axis. These treatments take several months to show urinary flow rate benefits but perhaps are ineffective in targeting androgen-independent basal cell proliferation. Surgical intervention by transurethral resection relieves urinary flow obstruction.
In normal somatic cells, repeated cell divisions result in an erosion of the repetitive terminal DNA sequences, the telomeres. The ultimate result of extreme telomere shortening is stress signal–mediated apoptosis or senescence. An understanding of telomerase biology is crucial to comprehend the proliferative dynamics of a tissue. Increased telomerase can be detected in approximately 70–90% of human cancers, including prostate cancer [2]. Although BPH tissue also exhibits a proliferative index two to three times higher than normal prostate [3], telomerase remains undetectable or is only sporadically expressed [2]. The inability to detect telomerase in non–stem-like cells, including the differentiated basal and luminal cells (>95% of the lesion), could have been interpreted as absent or sporadic telomerase expression or activity patterns in previous whole-biopsy analyses [2].
We assessed expression of the RNA (TERC) and protein (TERT) components of telomerase enzyme, telomerase activity, and telomere length in uncultured human BPH-derived stem-like cells (SCs: Lin−/CD31−/EpCAM+/CD133+/CD44+/α2β1hi), transit amplifying cells (TAs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1hi), committed basal cells (CBs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1lo), luminal cells (LCs: Lin−/CD31−/EpCAM+/CD44−/CD24+), and stromal cells (Lin−/CD31−/EpCAM−) (Supplementary Fig. 1, Supplement 1) [4] and [5] freshly purified from BPH tissues (Supplementary Table 1). The expression of NANOG was also higher in SCs (Supplementary Fig. 2). Previously, we reported that TERT expression was undetectable in normal prostate epithelial SCs and TAs [6]. In human BPH, however, the SCs and TAs exhibited significant overexpression of both TERC and TERT compared with the more differentiated CBs and LCs and stromal cells (Fig. 1a and 1b). Telomerase activity was also exclusively restricted to the SC/TA compartment, apart from one CB sample (Fig. 1c). All biopsies were specifically taken from the transitional zone with no detectable prostate cancer lesions on histology from patients with serum prostate-specific antigen levels <4 ng/ml to reduce the possibility of contamination with (telomerase-positive) cancer cells (Supplementary Table 1). TERT expression and telomerase activity were undetectable in stromal cells, indicating that telomerase is not essential for stromal hyperproliferation in BPH—another intriguing disparity in earlier data.

Epithelial cell telomere length measurements revealed that telomerase-expressing SCs and TAs had longer telomeres than those of their immediate differentiated progeny, the telomerase-negative CBs (p < 0.01) (Fig. 1d). The relative telomere length of LCs, however, was almost equivalent to that of SCs and TAs and was always longer than telomere lengths of patient-matched CBs (p < 0.01). Because LCs do not exhibit any detectable telomerase activity (Fig. 1c) and are believed to be derived from basal cells, the paradoxical observation of longer LC telomeres compared with CBs suggests that basal and luminal cells may be derived from distinct SC progenitors.
We stained BPH sections to determine basal cell content (using TP63) and luminal cell content (using NKX3.1) (Fig. 2a) and showed that basal and luminal hyperproliferation can occur together and even independently of each other because the basal:luminal ratio was highly variable, ranging from 0.22 to 2.08 (Fig. 2b, Supplementary Fig. 3), providing further evidence that basal and luminal cells may indeed be derived from different progenitors.

In human prostate, we have previously shown that SCs differentiate into CB cells via TA cells [4], unlike mouse cells, in which the situation is less clear [7]. The lineage of LCs, however, has not been determined in humans, although in vitro we can readily derive luminal cells from basal cells under specific culture conditions [5]. We now propose that basal and luminal cell lineages may develop separately in BPH and speculate that the prostate epithelial SCs can asymmetrically differentiate [8] into distinct basal and luminal progenitor cells, as shown in Figure 2c. In many other stem cell systems, the spindle orientation of progenitor progenies can determine their ultimate identity. The two-progenitor model may assist in understanding BPH pathogenesis, in which each progenitor and its progeny can expand independently or simultaneously to give rise to well-documented [9] areas of basal hyperproliferation, luminal hyperproliferation, or proliferation of both compartments, as shown in Figure 2a. If a basal progenitor cell differentiates into CBs, which retain limited proliferative potential in the absence of detectable telomerase activity, this will result in telomere shortening in CBs, for example, in basal cell hyperplasia, as shown in Figure 2a. In contrast, a luminal progenitor cell population expressing telomerase could expand and give rise to terminally differentiated luminal cells, which do not replicate, maintaining a telomere length similar to that of the luminal progenitors. Our data is in agreement with both Bonkhoff et al [10] and McNeal et al [11] with respect to the existence of a proliferative basal compartment in which the potential luminal progenitors derived from basal SCs exhibit characteristics of both basal and luminal phenotypes, analogous to castration-resistant Nkx3.1-expressing cells in mice [12]. The second implication of our model is that local inhibition of telomerase in BPH could be an alternative therapeutic strategy, as it will inhibit the proliferation both luminal and basal epithelial progenitors but perhaps not that of stromal cells. Because it is suggested that these progenitors function as a reservoir of proliferating epithelial cells in BPH, utilisation of novel and specific telomerase inhibitors such as Imetelstat (Geron Corp, Menlo Park, CA, USA) could prove to be a good adjuvant therapy for the management of BPH.
Author contributions: Norman J. Maitland had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Study concept and design: Maitland, Rane.
Acquisition of data: Rane, Frame, Greener.
Analysis and interpretation of data: Maitland, Rane, Frame.
Drafting of the manuscript: Rane.
Critical revision of the manuscript for important intellectual content: Maitland, Rane, Collins, Berney, Frame.
Statistical analysis: Rane, Frame.
Obtaining funding: Maitland.
Administrative, technical, or material support: Mann, Simms, Collins, Berney.
Supervision: Maitland.
Other (specify): None.
Financial disclosures: Norman J. Maitland certifies that all conflicts of interest, including specific financial interests and relationships and affiliations relevant to the subject matter or materials discussed in the manuscript (eg, employment/affiliation, grants or funding, consultancies, honoraria, stock ownership or options, expert testimony, royalties, or patents filed, received, or pending), are the following: None.
Funding/Support and role of the sponsor: The work was funded by Prostate Cancer UK (G2012-37) and PRO-NEST Marie-Curie network (J.K.R.), and Yorkshire Cancer Research (grant Y257PA, F.M.F. and N.J.M.). The human prostate tissue bank was funded by the Orchid Trust (D.B.). The sponsors were involved in the design and conduct of the study and collection and analysis of the data.
Acknowledgements: We would like to thank all of the patients and the urology surgeons L. Coombes, G. Cooksey and J. Hetherington (Castle Hill Hospital, Cottingham, UK). Thank you also to Megan Nash for immunofluorescence staining and counting and to Jerry Shay (University of Texas Southwestern, Dallas, TX, USA) for providing invaluable guidance on the telomerase and telomere measurement experiments. Davide Pellacani provided many useful discussions concerning the work.
Morbidity and loss of quality of life due to benign prostatic hyperplasia (BPH) are relatively high. There are >210 million cases of BPH worldwide [1], and the histologic prevalence of BPH is approximately 80% at the age of 80 yr. BPH is characterised by uncontrolled but noninvasive growth of a variety of cell types in the prostate, ranging in frequency from adenomyofibromatous, fibromuscular, fibroadenomatous, stromal, to rare muscular compositions. Medical treatments for BPH include targeting of the smooth muscle cells by α1-adrenergic receptor inhibitor and use of steroid 5α-reductase inhibitors to target the androgen-signalling axis. These treatments take several months to show urinary flow rate benefits but perhaps are ineffective in targeting androgen-independent basal cell proliferation. Surgical intervention by transurethral resection relieves urinary flow obstruction.
In normal somatic cells, repeated cell divisions result in an erosion of the repetitive terminal DNA sequences, the telomeres. The ultimate result of extreme telomere shortening is stress signal–mediated apoptosis or senescence. An understanding of telomerase biology is crucial to comprehend the proliferative dynamics of a tissue. Increased telomerase can be detected in approximately 70–90% of human cancers, including prostate cancer [2]. Although BPH tissue also exhibits a proliferative index two to three times higher than normal prostate [3], telomerase remains undetectable or is only sporadically expressed [2]. The inability to detect telomerase in non–stem-like cells, including the differentiated basal and luminal cells (>95% of the lesion), could have been interpreted as absent or sporadic telomerase expression or activity patterns in previous whole-biopsy analyses [2].
We assessed expression of the RNA (TERC) and protein (TERT) components of telomerase enzyme, telomerase activity, and telomere length in uncultured human BPH-derived stem-like cells (SCs: Lin−/CD31−/EpCAM+/CD133+/CD44+/α2β1hi), transit amplifying cells (TAs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1hi), committed basal cells (CBs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1lo), luminal cells (LCs: Lin−/CD31−/EpCAM+/CD44−/CD24+), and stromal cells (Lin−/CD31−/EpCAM−) (Supplementary Fig. 1, Supplement 1) [4] and [5] freshly purified from BPH tissues (Supplementary Table 1). The expression of NANOG was also higher in SCs (Supplementary Fig. 2). Previously, we reported that TERT expression was undetectable in normal prostate epithelial SCs and TAs [6]. In human BPH, however, the SCs and TAs exhibited significant overexpression of both TERC and TERT compared with the more differentiated CBs and LCs and stromal cells (Fig. 1a and 1b). Telomerase activity was also exclusively restricted to the SC/TA compartment, apart from one CB sample (Fig. 1c). All biopsies were specifically taken from the transitional zone with no detectable prostate cancer lesions on histology from patients with serum prostate-specific antigen levels <4 ng/ml to reduce the possibility of contamination with (telomerase-positive) cancer cells (Supplementary Table 1). TERT expression and telomerase activity were undetectable in stromal cells, indicating that telomerase is not essential for stromal hyperproliferation in BPH—another intriguing disparity in earlier data.

Epithelial cell telomere length measurements revealed that telomerase-expressing SCs and TAs had longer telomeres than those of their immediate differentiated progeny, the telomerase-negative CBs (p < 0.01) (Fig. 1d). The relative telomere length of LCs, however, was almost equivalent to that of SCs and TAs and was always longer than telomere lengths of patient-matched CBs (p < 0.01). Because LCs do not exhibit any detectable telomerase activity (Fig. 1c) and are believed to be derived from basal cells, the paradoxical observation of longer LC telomeres compared with CBs suggests that basal and luminal cells may be derived from distinct SC progenitors.
We stained BPH sections to determine basal cell content (using TP63) and luminal cell content (using NKX3.1) (Fig. 2a) and showed that basal and luminal hyperproliferation can occur together and even independently of each other because the basal:luminal ratio was highly variable, ranging from 0.22 to 2.08 (Fig. 2b, Supplementary Fig. 3), providing further evidence that basal and luminal cells may indeed be derived from different progenitors.

In human prostate, we have previously shown that SCs differentiate into CB cells via TA cells [4], unlike mouse cells, in which the situation is less clear [7]. The lineage of LCs, however, has not been determined in humans, although in vitro we can readily derive luminal cells from basal cells under specific culture conditions [5]. We now propose that basal and luminal cell lineages may develop separately in BPH and speculate that the prostate epithelial SCs can asymmetrically differentiate [8] into distinct basal and luminal progenitor cells, as shown in Figure 2c. In many other stem cell systems, the spindle orientation of progenitor progenies can determine their ultimate identity. The two-progenitor model may assist in understanding BPH pathogenesis, in which each progenitor and its progeny can expand independently or simultaneously to give rise to well-documented [9] areas of basal hyperproliferation, luminal hyperproliferation, or proliferation of both compartments, as shown in Figure 2a. If a basal progenitor cell differentiates into CBs, which retain limited proliferative potential in the absence of detectable telomerase activity, this will result in telomere shortening in CBs, for example, in basal cell hyperplasia, as shown in Figure 2a. In contrast, a luminal progenitor cell population expressing telomerase could expand and give rise to terminally differentiated luminal cells, which do not replicate, maintaining a telomere length similar to that of the luminal progenitors. Our data is in agreement with both Bonkhoff et al [10] and McNeal et al [11] with respect to the existence of a proliferative basal compartment in which the potential luminal progenitors derived from basal SCs exhibit characteristics of both basal and luminal phenotypes, analogous to castration-resistant Nkx3.1-expressing cells in mice [12]. The second implication of our model is that local inhibition of telomerase in BPH could be an alternative therapeutic strategy, as it will inhibit the proliferation both luminal and basal epithelial progenitors but perhaps not that of stromal cells. Because it is suggested that these progenitors function as a reservoir of proliferating epithelial cells in BPH, utilisation of novel and specific telomerase inhibitors such as Imetelstat (Geron Corp, Menlo Park, CA, USA) could prove to be a good adjuvant therapy for the management of BPH.
Author contributions: Norman J. Maitland had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Study concept and design: Maitland, Rane.
Acquisition of data: Rane, Frame, Greener.
Analysis and interpretation of data: Maitland, Rane, Frame.
Drafting of the manuscript: Rane.
Critical revision of the manuscript for important intellectual content: Maitland, Rane, Collins, Berney, Frame.
Statistical analysis: Rane, Frame.
Obtaining funding: Maitland.
Administrative, technical, or material support: Mann, Simms, Collins, Berney.
Supervision: Maitland.
Other (specify): None.
Financial disclosures: Norman J. Maitland certifies that all conflicts of interest, including specific financial interests and relationships and affiliations relevant to the subject matter or materials discussed in the manuscript (eg, employment/affiliation, grants or funding, consultancies, honoraria, stock ownership or options, expert testimony, royalties, or patents filed, received, or pending), are the following: None.
Funding/Support and role of the sponsor: The work was funded by Prostate Cancer UK (G2012-37) and PRO-NEST Marie-Curie network (J.K.R.), and Yorkshire Cancer Research (grant Y257PA, F.M.F. and N.J.M.). The human prostate tissue bank was funded by the Orchid Trust (D.B.). The sponsors were involved in the design and conduct of the study and collection and analysis of the data.
Acknowledgements: We would like to thank all of the patients and the urology surgeons L. Coombes, G. Cooksey and J. Hetherington (Castle Hill Hospital, Cottingham, UK). Thank you also to Megan Nash for immunofluorescence staining and counting and to Jerry Shay (University of Texas Southwestern, Dallas, TX, USA) for providing invaluable guidance on the telomerase and telomere measurement experiments. Davide Pellacani provided many useful discussions concerning the work.
Morbidity and loss of quality of life due to benign prostatic hyperplasia (BPH) are relatively high. There are >210 million cases of BPH worldwide [1], and the histologic prevalence of BPH is approximately 80% at the age of 80 yr. BPH is characterised by uncontrolled but noninvasive growth of a variety of cell types in the prostate, ranging in frequency from adenomyofibromatous, fibromuscular, fibroadenomatous, stromal, to rare muscular compositions. Medical treatments for BPH include targeting of the smooth muscle cells by α1-adrenergic receptor inhibitor and use of steroid 5α-reductase inhibitors to target the androgen-signalling axis. These treatments take several months to show urinary flow rate benefits but perhaps are ineffective in targeting androgen-independent basal cell proliferation. Surgical intervention by transurethral resection relieves urinary flow obstruction.
In normal somatic cells, repeated cell divisions result in an erosion of the repetitive terminal DNA sequences, the telomeres. The ultimate result of extreme telomere shortening is stress signal–mediated apoptosis or senescence. An understanding of telomerase biology is crucial to comprehend the proliferative dynamics of a tissue. Increased telomerase can be detected in approximately 70–90% of human cancers, including prostate cancer [2]. Although BPH tissue also exhibits a proliferative index two to three times higher than normal prostate [3], telomerase remains undetectable or is only sporadically expressed [2]. The inability to detect telomerase in non–stem-like cells, including the differentiated basal and luminal cells (>95% of the lesion), could have been interpreted as absent or sporadic telomerase expression or activity patterns in previous whole-biopsy analyses [2].
We assessed expression of the RNA (TERC) and protein (TERT) components of telomerase enzyme, telomerase activity, and telomere length in uncultured human BPH-derived stem-like cells (SCs: Lin−/CD31−/EpCAM+/CD133+/CD44+/α2β1hi), transit amplifying cells (TAs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1hi), committed basal cells (CBs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1lo), luminal cells (LCs: Lin−/CD31−/EpCAM+/CD44−/CD24+), and stromal cells (Lin−/CD31−/EpCAM−) (Supplementary Fig. 1, Supplement 1) [4] and [5] freshly purified from BPH tissues (Supplementary Table 1). The expression of NANOG was also higher in SCs (Supplementary Fig. 2). Previously, we reported that TERT expression was undetectable in normal prostate epithelial SCs and TAs [6]. In human BPH, however, the SCs and TAs exhibited significant overexpression of both TERC and TERT compared with the more differentiated CBs and LCs and stromal cells (Fig. 1a and 1b). Telomerase activity was also exclusively restricted to the SC/TA compartment, apart from one CB sample (Fig. 1c). All biopsies were specifically taken from the transitional zone with no detectable prostate cancer lesions on histology from patients with serum prostate-specific antigen levels <4 ng/ml to reduce the possibility of contamination with (telomerase-positive) cancer cells (Supplementary Table 1). TERT expression and telomerase activity were undetectable in stromal cells, indicating that telomerase is not essential for stromal hyperproliferation in BPH—another intriguing disparity in earlier data.

Epithelial cell telomere length measurements revealed that telomerase-expressing SCs and TAs had longer telomeres than those of their immediate differentiated progeny, the telomerase-negative CBs (p < 0.01) (Fig. 1d). The relative telomere length of LCs, however, was almost equivalent to that of SCs and TAs and was always longer than telomere lengths of patient-matched CBs (p < 0.01). Because LCs do not exhibit any detectable telomerase activity (Fig. 1c) and are believed to be derived from basal cells, the paradoxical observation of longer LC telomeres compared with CBs suggests that basal and luminal cells may be derived from distinct SC progenitors.
We stained BPH sections to determine basal cell content (using TP63) and luminal cell content (using NKX3.1) (Fig. 2a) and showed that basal and luminal hyperproliferation can occur together and even independently of each other because the basal:luminal ratio was highly variable, ranging from 0.22 to 2.08 (Fig. 2b, Supplementary Fig. 3), providing further evidence that basal and luminal cells may indeed be derived from different progenitors.

In human prostate, we have previously shown that SCs differentiate into CB cells via TA cells [4], unlike mouse cells, in which the situation is less clear [7]. The lineage of LCs, however, has not been determined in humans, although in vitro we can readily derive luminal cells from basal cells under specific culture conditions [5]. We now propose that basal and luminal cell lineages may develop separately in BPH and speculate that the prostate epithelial SCs can asymmetrically differentiate [8] into distinct basal and luminal progenitor cells, as shown in Figure 2c. In many other stem cell systems, the spindle orientation of progenitor progenies can determine their ultimate identity. The two-progenitor model may assist in understanding BPH pathogenesis, in which each progenitor and its progeny can expand independently or simultaneously to give rise to well-documented [9] areas of basal hyperproliferation, luminal hyperproliferation, or proliferation of both compartments, as shown in Figure 2a. If a basal progenitor cell differentiates into CBs, which retain limited proliferative potential in the absence of detectable telomerase activity, this will result in telomere shortening in CBs, for example, in basal cell hyperplasia, as shown in Figure 2a. In contrast, a luminal progenitor cell population expressing telomerase could expand and give rise to terminally differentiated luminal cells, which do not replicate, maintaining a telomere length similar to that of the luminal progenitors. Our data is in agreement with both Bonkhoff et al [10] and McNeal et al [11] with respect to the existence of a proliferative basal compartment in which the potential luminal progenitors derived from basal SCs exhibit characteristics of both basal and luminal phenotypes, analogous to castration-resistant Nkx3.1-expressing cells in mice [12]. The second implication of our model is that local inhibition of telomerase in BPH could be an alternative therapeutic strategy, as it will inhibit the proliferation both luminal and basal epithelial progenitors but perhaps not that of stromal cells. Because it is suggested that these progenitors function as a reservoir of proliferating epithelial cells in BPH, utilisation of novel and specific telomerase inhibitors such as Imetelstat (Geron Corp, Menlo Park, CA, USA) could prove to be a good adjuvant therapy for the management of BPH.
Author contributions: Norman J. Maitland had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Study concept and design: Maitland, Rane.
Acquisition of data: Rane, Frame, Greener.
Analysis and interpretation of data: Maitland, Rane, Frame.
Drafting of the manuscript: Rane.
Critical revision of the manuscript for important intellectual content: Maitland, Rane, Collins, Berney, Frame.
Statistical analysis: Rane, Frame.
Obtaining funding: Maitland.
Administrative, technical, or material support: Mann, Simms, Collins, Berney.
Supervision: Maitland.
Other (specify): None.
Financial disclosures: Norman J. Maitland certifies that all conflicts of interest, including specific financial interests and relationships and affiliations relevant to the subject matter or materials discussed in the manuscript (eg, employment/affiliation, grants or funding, consultancies, honoraria, stock ownership or options, expert testimony, royalties, or patents filed, received, or pending), are the following: None.
Funding/Support and role of the sponsor: The work was funded by Prostate Cancer UK (G2012-37) and PRO-NEST Marie-Curie network (J.K.R.), and Yorkshire Cancer Research (grant Y257PA, F.M.F. and N.J.M.). The human prostate tissue bank was funded by the Orchid Trust (D.B.). The sponsors were involved in the design and conduct of the study and collection and analysis of the data.
Acknowledgements: We would like to thank all of the patients and the urology surgeons L. Coombes, G. Cooksey and J. Hetherington (Castle Hill Hospital, Cottingham, UK). Thank you also to Megan Nash for immunofluorescence staining and counting and to Jerry Shay (University of Texas Southwestern, Dallas, TX, USA) for providing invaluable guidance on the telomerase and telomere measurement experiments. Davide Pellacani provided many useful discussions concerning the work.
Morbidity and loss of quality of life due to benign prostatic hyperplasia (BPH) are relatively high. There are >210 million cases of BPH worldwide [1], and the histologic prevalence of BPH is approximately 80% at the age of 80 yr. BPH is characterised by uncontrolled but noninvasive growth of a variety of cell types in the prostate, ranging in frequency from adenomyofibromatous, fibromuscular, fibroadenomatous, stromal, to rare muscular compositions. Medical treatments for BPH include targeting of the smooth muscle cells by α1-adrenergic receptor inhibitor and use of steroid 5α-reductase inhibitors to target the androgen-signalling axis. These treatments take several months to show urinary flow rate benefits but perhaps are ineffective in targeting androgen-independent basal cell proliferation. Surgical intervention by transurethral resection relieves urinary flow obstruction.
In normal somatic cells, repeated cell divisions result in an erosion of the repetitive terminal DNA sequences, the telomeres. The ultimate result of extreme telomere shortening is stress signal–mediated apoptosis or senescence. An understanding of telomerase biology is crucial to comprehend the proliferative dynamics of a tissue. Increased telomerase can be detected in approximately 70–90% of human cancers, including prostate cancer [2]. Although BPH tissue also exhibits a proliferative index two to three times higher than normal prostate [3], telomerase remains undetectable or is only sporadically expressed [2]. The inability to detect telomerase in non–stem-like cells, including the differentiated basal and luminal cells (>95% of the lesion), could have been interpreted as absent or sporadic telomerase expression or activity patterns in previous whole-biopsy analyses [2].
We assessed expression of the RNA (TERC) and protein (TERT) components of telomerase enzyme, telomerase activity, and telomere length in uncultured human BPH-derived stem-like cells (SCs: Lin−/CD31−/EpCAM+/CD133+/CD44+/α2β1hi), transit amplifying cells (TAs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1hi), committed basal cells (CBs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1lo), luminal cells (LCs: Lin−/CD31−/EpCAM+/CD44−/CD24+), and stromal cells (Lin−/CD31−/EpCAM−) (Supplementary Fig. 1, Supplement 1) [4] and [5] freshly purified from BPH tissues (Supplementary Table 1). The expression of NANOG was also higher in SCs (Supplementary Fig. 2). Previously, we reported that TERT expression was undetectable in normal prostate epithelial SCs and TAs [6]. In human BPH, however, the SCs and TAs exhibited significant overexpression of both TERC and TERT compared with the more differentiated CBs and LCs and stromal cells (Fig. 1a and 1b). Telomerase activity was also exclusively restricted to the SC/TA compartment, apart from one CB sample (Fig. 1c). All biopsies were specifically taken from the transitional zone with no detectable prostate cancer lesions on histology from patients with serum prostate-specific antigen levels <4 ng/ml to reduce the possibility of contamination with (telomerase-positive) cancer cells (Supplementary Table 1). TERT expression and telomerase activity were undetectable in stromal cells, indicating that telomerase is not essential for stromal hyperproliferation in BPH—another intriguing disparity in earlier data.

Epithelial cell telomere length measurements revealed that telomerase-expressing SCs and TAs had longer telomeres than those of their immediate differentiated progeny, the telomerase-negative CBs (p < 0.01) (Fig. 1d). The relative telomere length of LCs, however, was almost equivalent to that of SCs and TAs and was always longer than telomere lengths of patient-matched CBs (p < 0.01). Because LCs do not exhibit any detectable telomerase activity (Fig. 1c) and are believed to be derived from basal cells, the paradoxical observation of longer LC telomeres compared with CBs suggests that basal and luminal cells may be derived from distinct SC progenitors.
We stained BPH sections to determine basal cell content (using TP63) and luminal cell content (using NKX3.1) (Fig. 2a) and showed that basal and luminal hyperproliferation can occur together and even independently of each other because the basal:luminal ratio was highly variable, ranging from 0.22 to 2.08 (Fig. 2b, Supplementary Fig. 3), providing further evidence that basal and luminal cells may indeed be derived from different progenitors.

In human prostate, we have previously shown that SCs differentiate into CB cells via TA cells [4], unlike mouse cells, in which the situation is less clear [7]. The lineage of LCs, however, has not been determined in humans, although in vitro we can readily derive luminal cells from basal cells under specific culture conditions [5]. We now propose that basal and luminal cell lineages may develop separately in BPH and speculate that the prostate epithelial SCs can asymmetrically differentiate [8] into distinct basal and luminal progenitor cells, as shown in Figure 2c. In many other stem cell systems, the spindle orientation of progenitor progenies can determine their ultimate identity. The two-progenitor model may assist in understanding BPH pathogenesis, in which each progenitor and its progeny can expand independently or simultaneously to give rise to well-documented [9] areas of basal hyperproliferation, luminal hyperproliferation, or proliferation of both compartments, as shown in Figure 2a. If a basal progenitor cell differentiates into CBs, which retain limited proliferative potential in the absence of detectable telomerase activity, this will result in telomere shortening in CBs, for example, in basal cell hyperplasia, as shown in Figure 2a. In contrast, a luminal progenitor cell population expressing telomerase could expand and give rise to terminally differentiated luminal cells, which do not replicate, maintaining a telomere length similar to that of the luminal progenitors. Our data is in agreement with both Bonkhoff et al [10] and McNeal et al [11] with respect to the existence of a proliferative basal compartment in which the potential luminal progenitors derived from basal SCs exhibit characteristics of both basal and luminal phenotypes, analogous to castration-resistant Nkx3.1-expressing cells in mice [12]. The second implication of our model is that local inhibition of telomerase in BPH could be an alternative therapeutic strategy, as it will inhibit the proliferation both luminal and basal epithelial progenitors but perhaps not that of stromal cells. Because it is suggested that these progenitors function as a reservoir of proliferating epithelial cells in BPH, utilisation of novel and specific telomerase inhibitors such as Imetelstat (Geron Corp, Menlo Park, CA, USA) could prove to be a good adjuvant therapy for the management of BPH.
Author contributions: Norman J. Maitland had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Study concept and design: Maitland, Rane.
Acquisition of data: Rane, Frame, Greener.
Analysis and interpretation of data: Maitland, Rane, Frame.
Drafting of the manuscript: Rane.
Critical revision of the manuscript for important intellectual content: Maitland, Rane, Collins, Berney, Frame.
Statistical analysis: Rane, Frame.
Obtaining funding: Maitland.
Administrative, technical, or material support: Mann, Simms, Collins, Berney.
Supervision: Maitland.
Other (specify): None.
Financial disclosures: Norman J. Maitland certifies that all conflicts of interest, including specific financial interests and relationships and affiliations relevant to the subject matter or materials discussed in the manuscript (eg, employment/affiliation, grants or funding, consultancies, honoraria, stock ownership or options, expert testimony, royalties, or patents filed, received, or pending), are the following: None.
Funding/Support and role of the sponsor: The work was funded by Prostate Cancer UK (G2012-37) and PRO-NEST Marie-Curie network (J.K.R.), and Yorkshire Cancer Research (grant Y257PA, F.M.F. and N.J.M.). The human prostate tissue bank was funded by the Orchid Trust (D.B.). The sponsors were involved in the design and conduct of the study and collection and analysis of the data.
Acknowledgements: We would like to thank all of the patients and the urology surgeons L. Coombes, G. Cooksey and J. Hetherington (Castle Hill Hospital, Cottingham, UK). Thank you also to Megan Nash for immunofluorescence staining and counting and to Jerry Shay (University of Texas Southwestern, Dallas, TX, USA) for providing invaluable guidance on the telomerase and telomere measurement experiments. Davide Pellacani provided many useful discussions concerning the work.
Morbidity and loss of quality of life due to benign prostatic hyperplasia (BPH) are relatively high. There are >210 million cases of BPH worldwide [1], and the histologic prevalence of BPH is approximately 80% at the age of 80 yr. BPH is characterised by uncontrolled but noninvasive growth of a variety of cell types in the prostate, ranging in frequency from adenomyofibromatous, fibromuscular, fibroadenomatous, stromal, to rare muscular compositions. Medical treatments for BPH include targeting of the smooth muscle cells by α1-adrenergic receptor inhibitor and use of steroid 5α-reductase inhibitors to target the androgen-signalling axis. These treatments take several months to show urinary flow rate benefits but perhaps are ineffective in targeting androgen-independent basal cell proliferation. Surgical intervention by transurethral resection relieves urinary flow obstruction.
In normal somatic cells, repeated cell divisions result in an erosion of the repetitive terminal DNA sequences, the telomeres. The ultimate result of extreme telomere shortening is stress signal–mediated apoptosis or senescence. An understanding of telomerase biology is crucial to comprehend the proliferative dynamics of a tissue. Increased telomerase can be detected in approximately 70–90% of human cancers, including prostate cancer [2]. Although BPH tissue also exhibits a proliferative index two to three times higher than normal prostate [3], telomerase remains undetectable or is only sporadically expressed [2]. The inability to detect telomerase in non–stem-like cells, including the differentiated basal and luminal cells (>95% of the lesion), could have been interpreted as absent or sporadic telomerase expression or activity patterns in previous whole-biopsy analyses [2].
We assessed expression of the RNA (TERC) and protein (TERT) components of telomerase enzyme, telomerase activity, and telomere length in uncultured human BPH-derived stem-like cells (SCs: Lin−/CD31−/EpCAM+/CD133+/CD44+/α2β1hi), transit amplifying cells (TAs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1hi), committed basal cells (CBs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1lo), luminal cells (LCs: Lin−/CD31−/EpCAM+/CD44−/CD24+), and stromal cells (Lin−/CD31−/EpCAM−) (Supplementary Fig. 1, Supplement 1) [4] and [5] freshly purified from BPH tissues (Supplementary Table 1). The expression of NANOG was also higher in SCs (Supplementary Fig. 2). Previously, we reported that TERT expression was undetectable in normal prostate epithelial SCs and TAs [6]. In human BPH, however, the SCs and TAs exhibited significant overexpression of both TERC and TERT compared with the more differentiated CBs and LCs and stromal cells (Fig. 1a and 1b). Telomerase activity was also exclusively restricted to the SC/TA compartment, apart from one CB sample (Fig. 1c). All biopsies were specifically taken from the transitional zone with no detectable prostate cancer lesions on histology from patients with serum prostate-specific antigen levels <4 ng/ml to reduce the possibility of contamination with (telomerase-positive) cancer cells (Supplementary Table 1). TERT expression and telomerase activity were undetectable in stromal cells, indicating that telomerase is not essential for stromal hyperproliferation in BPH—another intriguing disparity in earlier data.

Epithelial cell telomere length measurements revealed that telomerase-expressing SCs and TAs had longer telomeres than those of their immediate differentiated progeny, the telomerase-negative CBs (p < 0.01) (Fig. 1d). The relative telomere length of LCs, however, was almost equivalent to that of SCs and TAs and was always longer than telomere lengths of patient-matched CBs (p < 0.01). Because LCs do not exhibit any detectable telomerase activity (Fig. 1c) and are believed to be derived from basal cells, the paradoxical observation of longer LC telomeres compared with CBs suggests that basal and luminal cells may be derived from distinct SC progenitors.
We stained BPH sections to determine basal cell content (using TP63) and luminal cell content (using NKX3.1) (Fig. 2a) and showed that basal and luminal hyperproliferation can occur together and even independently of each other because the basal:luminal ratio was highly variable, ranging from 0.22 to 2.08 (Fig. 2b, Supplementary Fig. 3), providing further evidence that basal and luminal cells may indeed be derived from different progenitors.

In human prostate, we have previously shown that SCs differentiate into CB cells via TA cells [4], unlike mouse cells, in which the situation is less clear [7]. The lineage of LCs, however, has not been determined in humans, although in vitro we can readily derive luminal cells from basal cells under specific culture conditions [5]. We now propose that basal and luminal cell lineages may develop separately in BPH and speculate that the prostate epithelial SCs can asymmetrically differentiate [8] into distinct basal and luminal progenitor cells, as shown in Figure 2c. In many other stem cell systems, the spindle orientation of progenitor progenies can determine their ultimate identity. The two-progenitor model may assist in understanding BPH pathogenesis, in which each progenitor and its progeny can expand independently or simultaneously to give rise to well-documented [9] areas of basal hyperproliferation, luminal hyperproliferation, or proliferation of both compartments, as shown in Figure 2a. If a basal progenitor cell differentiates into CBs, which retain limited proliferative potential in the absence of detectable telomerase activity, this will result in telomere shortening in CBs, for example, in basal cell hyperplasia, as shown in Figure 2a. In contrast, a luminal progenitor cell population expressing telomerase could expand and give rise to terminally differentiated luminal cells, which do not replicate, maintaining a telomere length similar to that of the luminal progenitors. Our data is in agreement with both Bonkhoff et al [10] and McNeal et al [11] with respect to the existence of a proliferative basal compartment in which the potential luminal progenitors derived from basal SCs exhibit characteristics of both basal and luminal phenotypes, analogous to castration-resistant Nkx3.1-expressing cells in mice [12]. The second implication of our model is that local inhibition of telomerase in BPH could be an alternative therapeutic strategy, as it will inhibit the proliferation both luminal and basal epithelial progenitors but perhaps not that of stromal cells. Because it is suggested that these progenitors function as a reservoir of proliferating epithelial cells in BPH, utilisation of novel and specific telomerase inhibitors such as Imetelstat (Geron Corp, Menlo Park, CA, USA) could prove to be a good adjuvant therapy for the management of BPH.
Author contributions: Norman J. Maitland had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Study concept and design: Maitland, Rane.
Acquisition of data: Rane, Frame, Greener.
Analysis and interpretation of data: Maitland, Rane, Frame.
Drafting of the manuscript: Rane.
Critical revision of the manuscript for important intellectual content: Maitland, Rane, Collins, Berney, Frame.
Statistical analysis: Rane, Frame.
Obtaining funding: Maitland.
Administrative, technical, or material support: Mann, Simms, Collins, Berney.
Supervision: Maitland.
Other (specify): None.
Financial disclosures: Norman J. Maitland certifies that all conflicts of interest, including specific financial interests and relationships and affiliations relevant to the subject matter or materials discussed in the manuscript (eg, employment/affiliation, grants or funding, consultancies, honoraria, stock ownership or options, expert testimony, royalties, or patents filed, received, or pending), are the following: None.
Funding/Support and role of the sponsor: The work was funded by Prostate Cancer UK (G2012-37) and PRO-NEST Marie-Curie network (J.K.R.), and Yorkshire Cancer Research (grant Y257PA, F.M.F. and N.J.M.). The human prostate tissue bank was funded by the Orchid Trust (D.B.). The sponsors were involved in the design and conduct of the study and collection and analysis of the data.
Acknowledgements: We would like to thank all of the patients and the urology surgeons L. Coombes, G. Cooksey and J. Hetherington (Castle Hill Hospital, Cottingham, UK). Thank you also to Megan Nash for immunofluorescence staining and counting and to Jerry Shay (University of Texas Southwestern, Dallas, TX, USA) for providing invaluable guidance on the telomerase and telomere measurement experiments. Davide Pellacani provided many useful discussions concerning the work.
Morbidity and loss of quality of life due to benign prostatic hyperplasia (BPH) are relatively high. There are >210 million cases of BPH worldwide [1], and the histologic prevalence of BPH is approximately 80% at the age of 80 yr. BPH is characterised by uncontrolled but noninvasive growth of a variety of cell types in the prostate, ranging in frequency from adenomyofibromatous, fibromuscular, fibroadenomatous, stromal, to rare muscular compositions. Medical treatments for BPH include targeting of the smooth muscle cells by α1-adrenergic receptor inhibitor and use of steroid 5α-reductase inhibitors to target the androgen-signalling axis. These treatments take several months to show urinary flow rate benefits but perhaps are ineffective in targeting androgen-independent basal cell proliferation. Surgical intervention by transurethral resection relieves urinary flow obstruction.
In normal somatic cells, repeated cell divisions result in an erosion of the repetitive terminal DNA sequences, the telomeres. The ultimate result of extreme telomere shortening is stress signal–mediated apoptosis or senescence. An understanding of telomerase biology is crucial to comprehend the proliferative dynamics of a tissue. Increased telomerase can be detected in approximately 70–90% of human cancers, including prostate cancer [2]. Although BPH tissue also exhibits a proliferative index two to three times higher than normal prostate [3], telomerase remains undetectable or is only sporadically expressed [2]. The inability to detect telomerase in non–stem-like cells, including the differentiated basal and luminal cells (>95% of the lesion), could have been interpreted as absent or sporadic telomerase expression or activity patterns in previous whole-biopsy analyses [2].
We assessed expression of the RNA (TERC) and protein (TERT) components of telomerase enzyme, telomerase activity, and telomere length in uncultured human BPH-derived stem-like cells (SCs: Lin−/CD31−/EpCAM+/CD133+/CD44+/α2β1hi), transit amplifying cells (TAs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1hi), committed basal cells (CBs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1lo), luminal cells (LCs: Lin−/CD31−/EpCAM+/CD44−/CD24+), and stromal cells (Lin−/CD31−/EpCAM−) (Supplementary Fig. 1, Supplement 1) [4] and [5] freshly purified from BPH tissues (Supplementary Table 1). The expression of NANOG was also higher in SCs (Supplementary Fig. 2). Previously, we reported that TERT expression was undetectable in normal prostate epithelial SCs and TAs [6]. In human BPH, however, the SCs and TAs exhibited significant overexpression of both TERC and TERT compared with the more differentiated CBs and LCs and stromal cells (Fig. 1a and 1b). Telomerase activity was also exclusively restricted to the SC/TA compartment, apart from one CB sample (Fig. 1c). All biopsies were specifically taken from the transitional zone with no detectable prostate cancer lesions on histology from patients with serum prostate-specific antigen levels <4 ng/ml to reduce the possibility of contamination with (telomerase-positive) cancer cells (Supplementary Table 1). TERT expression and telomerase activity were undetectable in stromal cells, indicating that telomerase is not essential for stromal hyperproliferation in BPH—another intriguing disparity in earlier data.

Epithelial cell telomere length measurements revealed that telomerase-expressing SCs and TAs had longer telomeres than those of their immediate differentiated progeny, the telomerase-negative CBs (p < 0.01) (Fig. 1d). The relative telomere length of LCs, however, was almost equivalent to that of SCs and TAs and was always longer than telomere lengths of patient-matched CBs (p < 0.01). Because LCs do not exhibit any detectable telomerase activity (Fig. 1c) and are believed to be derived from basal cells, the paradoxical observation of longer LC telomeres compared with CBs suggests that basal and luminal cells may be derived from distinct SC progenitors.
We stained BPH sections to determine basal cell content (using TP63) and luminal cell content (using NKX3.1) (Fig. 2a) and showed that basal and luminal hyperproliferation can occur together and even independently of each other because the basal:luminal ratio was highly variable, ranging from 0.22 to 2.08 (Fig. 2b, Supplementary Fig. 3), providing further evidence that basal and luminal cells may indeed be derived from different progenitors.

In human prostate, we have previously shown that SCs differentiate into CB cells via TA cells [4], unlike mouse cells, in which the situation is less clear [7]. The lineage of LCs, however, has not been determined in humans, although in vitro we can readily derive luminal cells from basal cells under specific culture conditions [5]. We now propose that basal and luminal cell lineages may develop separately in BPH and speculate that the prostate epithelial SCs can asymmetrically differentiate [8] into distinct basal and luminal progenitor cells, as shown in Figure 2c. In many other stem cell systems, the spindle orientation of progenitor progenies can determine their ultimate identity. The two-progenitor model may assist in understanding BPH pathogenesis, in which each progenitor and its progeny can expand independently or simultaneously to give rise to well-documented [9] areas of basal hyperproliferation, luminal hyperproliferation, or proliferation of both compartments, as shown in Figure 2a. If a basal progenitor cell differentiates into CBs, which retain limited proliferative potential in the absence of detectable telomerase activity, this will result in telomere shortening in CBs, for example, in basal cell hyperplasia, as shown in Figure 2a. In contrast, a luminal progenitor cell population expressing telomerase could expand and give rise to terminally differentiated luminal cells, which do not replicate, maintaining a telomere length similar to that of the luminal progenitors. Our data is in agreement with both Bonkhoff et al [10] and McNeal et al [11] with respect to the existence of a proliferative basal compartment in which the potential luminal progenitors derived from basal SCs exhibit characteristics of both basal and luminal phenotypes, analogous to castration-resistant Nkx3.1-expressing cells in mice [12]. The second implication of our model is that local inhibition of telomerase in BPH could be an alternative therapeutic strategy, as it will inhibit the proliferation both luminal and basal epithelial progenitors but perhaps not that of stromal cells. Because it is suggested that these progenitors function as a reservoir of proliferating epithelial cells in BPH, utilisation of novel and specific telomerase inhibitors such as Imetelstat (Geron Corp, Menlo Park, CA, USA) could prove to be a good adjuvant therapy for the management of BPH.
Author contributions: Norman J. Maitland had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Study concept and design: Maitland, Rane.
Acquisition of data: Rane, Frame, Greener.
Analysis and interpretation of data: Maitland, Rane, Frame.
Drafting of the manuscript: Rane.
Critical revision of the manuscript for important intellectual content: Maitland, Rane, Collins, Berney, Frame.
Statistical analysis: Rane, Frame.
Obtaining funding: Maitland.
Administrative, technical, or material support: Mann, Simms, Collins, Berney.
Supervision: Maitland.
Other (specify): None.
Financial disclosures: Norman J. Maitland certifies that all conflicts of interest, including specific financial interests and relationships and affiliations relevant to the subject matter or materials discussed in the manuscript (eg, employment/affiliation, grants or funding, consultancies, honoraria, stock ownership or options, expert testimony, royalties, or patents filed, received, or pending), are the following: None.
Funding/Support and role of the sponsor: The work was funded by Prostate Cancer UK (G2012-37) and PRO-NEST Marie-Curie network (J.K.R.), and Yorkshire Cancer Research (grant Y257PA, F.M.F. and N.J.M.). The human prostate tissue bank was funded by the Orchid Trust (D.B.). The sponsors were involved in the design and conduct of the study and collection and analysis of the data.
Acknowledgements: We would like to thank all of the patients and the urology surgeons L. Coombes, G. Cooksey and J. Hetherington (Castle Hill Hospital, Cottingham, UK). Thank you also to Megan Nash for immunofluorescence staining and counting and to Jerry Shay (University of Texas Southwestern, Dallas, TX, USA) for providing invaluable guidance on the telomerase and telomere measurement experiments. Davide Pellacani provided many useful discussions concerning the work.
Morbidity and loss of quality of life due to benign prostatic hyperplasia (BPH) are relatively high. There are >210 million cases of BPH worldwide [1], and the histologic prevalence of BPH is approximately 80% at the age of 80 yr. BPH is characterised by uncontrolled but noninvasive growth of a variety of cell types in the prostate, ranging in frequency from adenomyofibromatous, fibromuscular, fibroadenomatous, stromal, to rare muscular compositions. Medical treatments for BPH include targeting of the smooth muscle cells by α1-adrenergic receptor inhibitor and use of steroid 5α-reductase inhibitors to target the androgen-signalling axis. These treatments take several months to show urinary flow rate benefits but perhaps are ineffective in targeting androgen-independent basal cell proliferation. Surgical intervention by transurethral resection relieves urinary flow obstruction.
In normal somatic cells, repeated cell divisions result in an erosion of the repetitive terminal DNA sequences, the telomeres. The ultimate result of extreme telomere shortening is stress signal–mediated apoptosis or senescence. An understanding of telomerase biology is crucial to comprehend the proliferative dynamics of a tissue. Increased telomerase can be detected in approximately 70–90% of human cancers, including prostate cancer [2]. Although BPH tissue also exhibits a proliferative index two to three times higher than normal prostate [3], telomerase remains undetectable or is only sporadically expressed [2]. The inability to detect telomerase in non–stem-like cells, including the differentiated basal and luminal cells (>95% of the lesion), could have been interpreted as absent or sporadic telomerase expression or activity patterns in previous whole-biopsy analyses [2].
We assessed expression of the RNA (TERC) and protein (TERT) components of telomerase enzyme, telomerase activity, and telomere length in uncultured human BPH-derived stem-like cells (SCs: Lin−/CD31−/EpCAM+/CD133+/CD44+/α2β1hi), transit amplifying cells (TAs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1hi), committed basal cells (CBs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1lo), luminal cells (LCs: Lin−/CD31−/EpCAM+/CD44−/CD24+), and stromal cells (Lin−/CD31−/EpCAM−) (Supplementary Fig. 1, Supplement 1) [4] and [5] freshly purified from BPH tissues (Supplementary Table 1). The expression of NANOG was also higher in SCs (Supplementary Fig. 2). Previously, we reported that TERT expression was undetectable in normal prostate epithelial SCs and TAs [6]. In human BPH, however, the SCs and TAs exhibited significant overexpression of both TERC and TERT compared with the more differentiated CBs and LCs and stromal cells (Fig. 1a and 1b). Telomerase activity was also exclusively restricted to the SC/TA compartment, apart from one CB sample (Fig. 1c). All biopsies were specifically taken from the transitional zone with no detectable prostate cancer lesions on histology from patients with serum prostate-specific antigen levels <4 ng/ml to reduce the possibility of contamination with (telomerase-positive) cancer cells (Supplementary Table 1). TERT expression and telomerase activity were undetectable in stromal cells, indicating that telomerase is not essential for stromal hyperproliferation in BPH—another intriguing disparity in earlier data.

Epithelial cell telomere length measurements revealed that telomerase-expressing SCs and TAs had longer telomeres than those of their immediate differentiated progeny, the telomerase-negative CBs (p < 0.01) (Fig. 1d). The relative telomere length of LCs, however, was almost equivalent to that of SCs and TAs and was always longer than telomere lengths of patient-matched CBs (p < 0.01). Because LCs do not exhibit any detectable telomerase activity (Fig. 1c) and are believed to be derived from basal cells, the paradoxical observation of longer LC telomeres compared with CBs suggests that basal and luminal cells may be derived from distinct SC progenitors.
We stained BPH sections to determine basal cell content (using TP63) and luminal cell content (using NKX3.1) (Fig. 2a) and showed that basal and luminal hyperproliferation can occur together and even independently of each other because the basal:luminal ratio was highly variable, ranging from 0.22 to 2.08 (Fig. 2b, Supplementary Fig. 3), providing further evidence that basal and luminal cells may indeed be derived from different progenitors.

In human prostate, we have previously shown that SCs differentiate into CB cells via TA cells [4], unlike mouse cells, in which the situation is less clear [7]. The lineage of LCs, however, has not been determined in humans, although in vitro we can readily derive luminal cells from basal cells under specific culture conditions [5]. We now propose that basal and luminal cell lineages may develop separately in BPH and speculate that the prostate epithelial SCs can asymmetrically differentiate [8] into distinct basal and luminal progenitor cells, as shown in Figure 2c. In many other stem cell systems, the spindle orientation of progenitor progenies can determine their ultimate identity. The two-progenitor model may assist in understanding BPH pathogenesis, in which each progenitor and its progeny can expand independently or simultaneously to give rise to well-documented [9] areas of basal hyperproliferation, luminal hyperproliferation, or proliferation of both compartments, as shown in Figure 2a. If a basal progenitor cell differentiates into CBs, which retain limited proliferative potential in the absence of detectable telomerase activity, this will result in telomere shortening in CBs, for example, in basal cell hyperplasia, as shown in Figure 2a. In contrast, a luminal progenitor cell population expressing telomerase could expand and give rise to terminally differentiated luminal cells, which do not replicate, maintaining a telomere length similar to that of the luminal progenitors. Our data is in agreement with both Bonkhoff et al [10] and McNeal et al [11] with respect to the existence of a proliferative basal compartment in which the potential luminal progenitors derived from basal SCs exhibit characteristics of both basal and luminal phenotypes, analogous to castration-resistant Nkx3.1-expressing cells in mice [12]. The second implication of our model is that local inhibition of telomerase in BPH could be an alternative therapeutic strategy, as it will inhibit the proliferation both luminal and basal epithelial progenitors but perhaps not that of stromal cells. Because it is suggested that these progenitors function as a reservoir of proliferating epithelial cells in BPH, utilisation of novel and specific telomerase inhibitors such as Imetelstat (Geron Corp, Menlo Park, CA, USA) could prove to be a good adjuvant therapy for the management of BPH.
Author contributions: Norman J. Maitland had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Study concept and design: Maitland, Rane.
Acquisition of data: Rane, Frame, Greener.
Analysis and interpretation of data: Maitland, Rane, Frame.
Drafting of the manuscript: Rane.
Critical revision of the manuscript for important intellectual content: Maitland, Rane, Collins, Berney, Frame.
Statistical analysis: Rane, Frame.
Obtaining funding: Maitland.
Administrative, technical, or material support: Mann, Simms, Collins, Berney.
Supervision: Maitland.
Other (specify): None.
Financial disclosures: Norman J. Maitland certifies that all conflicts of interest, including specific financial interests and relationships and affiliations relevant to the subject matter or materials discussed in the manuscript (eg, employment/affiliation, grants or funding, consultancies, honoraria, stock ownership or options, expert testimony, royalties, or patents filed, received, or pending), are the following: None.
Funding/Support and role of the sponsor: The work was funded by Prostate Cancer UK (G2012-37) and PRO-NEST Marie-Curie network (J.K.R.), and Yorkshire Cancer Research (grant Y257PA, F.M.F. and N.J.M.). The human prostate tissue bank was funded by the Orchid Trust (D.B.). The sponsors were involved in the design and conduct of the study and collection and analysis of the data.
Acknowledgements: We would like to thank all of the patients and the urology surgeons L. Coombes, G. Cooksey and J. Hetherington (Castle Hill Hospital, Cottingham, UK). Thank you also to Megan Nash for immunofluorescence staining and counting and to Jerry Shay (University of Texas Southwestern, Dallas, TX, USA) for providing invaluable guidance on the telomerase and telomere measurement experiments. Davide Pellacani provided many useful discussions concerning the work.
Morbidity and loss of quality of life due to benign prostatic hyperplasia (BPH) are relatively high. There are >210 million cases of BPH worldwide [1], and the histologic prevalence of BPH is approximately 80% at the age of 80 yr. BPH is characterised by uncontrolled but noninvasive growth of a variety of cell types in the prostate, ranging in frequency from adenomyofibromatous, fibromuscular, fibroadenomatous, stromal, to rare muscular compositions. Medical treatments for BPH include targeting of the smooth muscle cells by α1-adrenergic receptor inhibitor and use of steroid 5α-reductase inhibitors to target the androgen-signalling axis. These treatments take several months to show urinary flow rate benefits but perhaps are ineffective in targeting androgen-independent basal cell proliferation. Surgical intervention by transurethral resection relieves urinary flow obstruction.
In normal somatic cells, repeated cell divisions result in an erosion of the repetitive terminal DNA sequences, the telomeres. The ultimate result of extreme telomere shortening is stress signal–mediated apoptosis or senescence. An understanding of telomerase biology is crucial to comprehend the proliferative dynamics of a tissue. Increased telomerase can be detected in approximately 70–90% of human cancers, including prostate cancer [2]. Although BPH tissue also exhibits a proliferative index two to three times higher than normal prostate [3], telomerase remains undetectable or is only sporadically expressed [2]. The inability to detect telomerase in non–stem-like cells, including the differentiated basal and luminal cells (>95% of the lesion), could have been interpreted as absent or sporadic telomerase expression or activity patterns in previous whole-biopsy analyses [2].
We assessed expression of the RNA (TERC) and protein (TERT) components of telomerase enzyme, telomerase activity, and telomere length in uncultured human BPH-derived stem-like cells (SCs: Lin−/CD31−/EpCAM+/CD133+/CD44+/α2β1hi), transit amplifying cells (TAs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1hi), committed basal cells (CBs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1lo), luminal cells (LCs: Lin−/CD31−/EpCAM+/CD44−/CD24+), and stromal cells (Lin−/CD31−/EpCAM−) (Supplementary Fig. 1, Supplement 1) [4] and [5] freshly purified from BPH tissues (Supplementary Table 1). The expression of NANOG was also higher in SCs (Supplementary Fig. 2). Previously, we reported that TERT expression was undetectable in normal prostate epithelial SCs and TAs [6]. In human BPH, however, the SCs and TAs exhibited significant overexpression of both TERC and TERT compared with the more differentiated CBs and LCs and stromal cells (Fig. 1a and 1b). Telomerase activity was also exclusively restricted to the SC/TA compartment, apart from one CB sample (Fig. 1c). All biopsies were specifically taken from the transitional zone with no detectable prostate cancer lesions on histology from patients with serum prostate-specific antigen levels <4 ng/ml to reduce the possibility of contamination with (telomerase-positive) cancer cells (Supplementary Table 1). TERT expression and telomerase activity were undetectable in stromal cells, indicating that telomerase is not essential for stromal hyperproliferation in BPH—another intriguing disparity in earlier data.

Epithelial cell telomere length measurements revealed that telomerase-expressing SCs and TAs had longer telomeres than those of their immediate differentiated progeny, the telomerase-negative CBs (p < 0.01) (Fig. 1d). The relative telomere length of LCs, however, was almost equivalent to that of SCs and TAs and was always longer than telomere lengths of patient-matched CBs (p < 0.01). Because LCs do not exhibit any detectable telomerase activity (Fig. 1c) and are believed to be derived from basal cells, the paradoxical observation of longer LC telomeres compared with CBs suggests that basal and luminal cells may be derived from distinct SC progenitors.
We stained BPH sections to determine basal cell content (using TP63) and luminal cell content (using NKX3.1) (Fig. 2a) and showed that basal and luminal hyperproliferation can occur together and even independently of each other because the basal:luminal ratio was highly variable, ranging from 0.22 to 2.08 (Fig. 2b, Supplementary Fig. 3), providing further evidence that basal and luminal cells may indeed be derived from different progenitors.

In human prostate, we have previously shown that SCs differentiate into CB cells via TA cells [4], unlike mouse cells, in which the situation is less clear [7]. The lineage of LCs, however, has not been determined in humans, although in vitro we can readily derive luminal cells from basal cells under specific culture conditions [5]. We now propose that basal and luminal cell lineages may develop separately in BPH and speculate that the prostate epithelial SCs can asymmetrically differentiate [8] into distinct basal and luminal progenitor cells, as shown in Figure 2c. In many other stem cell systems, the spindle orientation of progenitor progenies can determine their ultimate identity. The two-progenitor model may assist in understanding BPH pathogenesis, in which each progenitor and its progeny can expand independently or simultaneously to give rise to well-documented [9] areas of basal hyperproliferation, luminal hyperproliferation, or proliferation of both compartments, as shown in Figure 2a. If a basal progenitor cell differentiates into CBs, which retain limited proliferative potential in the absence of detectable telomerase activity, this will result in telomere shortening in CBs, for example, in basal cell hyperplasia, as shown in Figure 2a. In contrast, a luminal progenitor cell population expressing telomerase could expand and give rise to terminally differentiated luminal cells, which do not replicate, maintaining a telomere length similar to that of the luminal progenitors. Our data is in agreement with both Bonkhoff et al [10] and McNeal et al [11] with respect to the existence of a proliferative basal compartment in which the potential luminal progenitors derived from basal SCs exhibit characteristics of both basal and luminal phenotypes, analogous to castration-resistant Nkx3.1-expressing cells in mice [12]. The second implication of our model is that local inhibition of telomerase in BPH could be an alternative therapeutic strategy, as it will inhibit the proliferation both luminal and basal epithelial progenitors but perhaps not that of stromal cells. Because it is suggested that these progenitors function as a reservoir of proliferating epithelial cells in BPH, utilisation of novel and specific telomerase inhibitors such as Imetelstat (Geron Corp, Menlo Park, CA, USA) could prove to be a good adjuvant therapy for the management of BPH.
Author contributions: Norman J. Maitland had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Study concept and design: Maitland, Rane.
Acquisition of data: Rane, Frame, Greener.
Analysis and interpretation of data: Maitland, Rane, Frame.
Drafting of the manuscript: Rane.
Critical revision of the manuscript for important intellectual content: Maitland, Rane, Collins, Berney, Frame.
Statistical analysis: Rane, Frame.
Obtaining funding: Maitland.
Administrative, technical, or material support: Mann, Simms, Collins, Berney.
Supervision: Maitland.
Other (specify): None.
Financial disclosures: Norman J. Maitland certifies that all conflicts of interest, including specific financial interests and relationships and affiliations relevant to the subject matter or materials discussed in the manuscript (eg, employment/affiliation, grants or funding, consultancies, honoraria, stock ownership or options, expert testimony, royalties, or patents filed, received, or pending), are the following: None.
Funding/Support and role of the sponsor: The work was funded by Prostate Cancer UK (G2012-37) and PRO-NEST Marie-Curie network (J.K.R.), and Yorkshire Cancer Research (grant Y257PA, F.M.F. and N.J.M.). The human prostate tissue bank was funded by the Orchid Trust (D.B.). The sponsors were involved in the design and conduct of the study and collection and analysis of the data.
Acknowledgements: We would like to thank all of the patients and the urology surgeons L. Coombes, G. Cooksey and J. Hetherington (Castle Hill Hospital, Cottingham, UK). Thank you also to Megan Nash for immunofluorescence staining and counting and to Jerry Shay (University of Texas Southwestern, Dallas, TX, USA) for providing invaluable guidance on the telomerase and telomere measurement experiments. Davide Pellacani provided many useful discussions concerning the work.
Morbidity and loss of quality of life due to benign prostatic hyperplasia (BPH) are relatively high. There are >210 million cases of BPH worldwide [1], and the histologic prevalence of BPH is approximately 80% at the age of 80 yr. BPH is characterised by uncontrolled but noninvasive growth of a variety of cell types in the prostate, ranging in frequency from adenomyofibromatous, fibromuscular, fibroadenomatous, stromal, to rare muscular compositions. Medical treatments for BPH include targeting of the smooth muscle cells by α1-adrenergic receptor inhibitor and use of steroid 5α-reductase inhibitors to target the androgen-signalling axis. These treatments take several months to show urinary flow rate benefits but perhaps are ineffective in targeting androgen-independent basal cell proliferation. Surgical intervention by transurethral resection relieves urinary flow obstruction.
In normal somatic cells, repeated cell divisions result in an erosion of the repetitive terminal DNA sequences, the telomeres. The ultimate result of extreme telomere shortening is stress signal–mediated apoptosis or senescence. An understanding of telomerase biology is crucial to comprehend the proliferative dynamics of a tissue. Increased telomerase can be detected in approximately 70–90% of human cancers, including prostate cancer [2]. Although BPH tissue also exhibits a proliferative index two to three times higher than normal prostate [3], telomerase remains undetectable or is only sporadically expressed [2]. The inability to detect telomerase in non–stem-like cells, including the differentiated basal and luminal cells (>95% of the lesion), could have been interpreted as absent or sporadic telomerase expression or activity patterns in previous whole-biopsy analyses [2].
We assessed expression of the RNA (TERC) and protein (TERT) components of telomerase enzyme, telomerase activity, and telomere length in uncultured human BPH-derived stem-like cells (SCs: Lin−/CD31−/EpCAM+/CD133+/CD44+/α2β1hi), transit amplifying cells (TAs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1hi), committed basal cells (CBs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1lo), luminal cells (LCs: Lin−/CD31−/EpCAM+/CD44−/CD24+), and stromal cells (Lin−/CD31−/EpCAM−) (Supplementary Fig. 1, Supplement 1) [4] and [5] freshly purified from BPH tissues (Supplementary Table 1). The expression of NANOG was also higher in SCs (Supplementary Fig. 2). Previously, we reported that TERT expression was undetectable in normal prostate epithelial SCs and TAs [6]. In human BPH, however, the SCs and TAs exhibited significant overexpression of both TERC and TERT compared with the more differentiated CBs and LCs and stromal cells (Fig. 1a and 1b). Telomerase activity was also exclusively restricted to the SC/TA compartment, apart from one CB sample (Fig. 1c). All biopsies were specifically taken from the transitional zone with no detectable prostate cancer lesions on histology from patients with serum prostate-specific antigen levels <4 ng/ml to reduce the possibility of contamination with (telomerase-positive) cancer cells (Supplementary Table 1). TERT expression and telomerase activity were undetectable in stromal cells, indicating that telomerase is not essential for stromal hyperproliferation in BPH—another intriguing disparity in earlier data.

Epithelial cell telomere length measurements revealed that telomerase-expressing SCs and TAs had longer telomeres than those of their immediate differentiated progeny, the telomerase-negative CBs (p < 0.01) (Fig. 1d). The relative telomere length of LCs, however, was almost equivalent to that of SCs and TAs and was always longer than telomere lengths of patient-matched CBs (p < 0.01). Because LCs do not exhibit any detectable telomerase activity (Fig. 1c) and are believed to be derived from basal cells, the paradoxical observation of longer LC telomeres compared with CBs suggests that basal and luminal cells may be derived from distinct SC progenitors.
We stained BPH sections to determine basal cell content (using TP63) and luminal cell content (using NKX3.1) (Fig. 2a) and showed that basal and luminal hyperproliferation can occur together and even independently of each other because the basal:luminal ratio was highly variable, ranging from 0.22 to 2.08 (Fig. 2b, Supplementary Fig. 3), providing further evidence that basal and luminal cells may indeed be derived from different progenitors.

In human prostate, we have previously shown that SCs differentiate into CB cells via TA cells [4], unlike mouse cells, in which the situation is less clear [7]. The lineage of LCs, however, has not been determined in humans, although in vitro we can readily derive luminal cells from basal cells under specific culture conditions [5]. We now propose that basal and luminal cell lineages may develop separately in BPH and speculate that the prostate epithelial SCs can asymmetrically differentiate [8] into distinct basal and luminal progenitor cells, as shown in Figure 2c. In many other stem cell systems, the spindle orientation of progenitor progenies can determine their ultimate identity. The two-progenitor model may assist in understanding BPH pathogenesis, in which each progenitor and its progeny can expand independently or simultaneously to give rise to well-documented [9] areas of basal hyperproliferation, luminal hyperproliferation, or proliferation of both compartments, as shown in Figure 2a. If a basal progenitor cell differentiates into CBs, which retain limited proliferative potential in the absence of detectable telomerase activity, this will result in telomere shortening in CBs, for example, in basal cell hyperplasia, as shown in Figure 2a. In contrast, a luminal progenitor cell population expressing telomerase could expand and give rise to terminally differentiated luminal cells, which do not replicate, maintaining a telomere length similar to that of the luminal progenitors. Our data is in agreement with both Bonkhoff et al [10] and McNeal et al [11] with respect to the existence of a proliferative basal compartment in which the potential luminal progenitors derived from basal SCs exhibit characteristics of both basal and luminal phenotypes, analogous to castration-resistant Nkx3.1-expressing cells in mice [12]. The second implication of our model is that local inhibition of telomerase in BPH could be an alternative therapeutic strategy, as it will inhibit the proliferation both luminal and basal epithelial progenitors but perhaps not that of stromal cells. Because it is suggested that these progenitors function as a reservoir of proliferating epithelial cells in BPH, utilisation of novel and specific telomerase inhibitors such as Imetelstat (Geron Corp, Menlo Park, CA, USA) could prove to be a good adjuvant therapy for the management of BPH.
Author contributions: Norman J. Maitland had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Study concept and design: Maitland, Rane.
Acquisition of data: Rane, Frame, Greener.
Analysis and interpretation of data: Maitland, Rane, Frame.
Drafting of the manuscript: Rane.
Critical revision of the manuscript for important intellectual content: Maitland, Rane, Collins, Berney, Frame.
Statistical analysis: Rane, Frame.
Obtaining funding: Maitland.
Administrative, technical, or material support: Mann, Simms, Collins, Berney.
Supervision: Maitland.
Other (specify): None.
Financial disclosures: Norman J. Maitland certifies that all conflicts of interest, including specific financial interests and relationships and affiliations relevant to the subject matter or materials discussed in the manuscript (eg, employment/affiliation, grants or funding, consultancies, honoraria, stock ownership or options, expert testimony, royalties, or patents filed, received, or pending), are the following: None.
Funding/Support and role of the sponsor: The work was funded by Prostate Cancer UK (G2012-37) and PRO-NEST Marie-Curie network (J.K.R.), and Yorkshire Cancer Research (grant Y257PA, F.M.F. and N.J.M.). The human prostate tissue bank was funded by the Orchid Trust (D.B.). The sponsors were involved in the design and conduct of the study and collection and analysis of the data.
Acknowledgements: We would like to thank all of the patients and the urology surgeons L. Coombes, G. Cooksey and J. Hetherington (Castle Hill Hospital, Cottingham, UK). Thank you also to Megan Nash for immunofluorescence staining and counting and to Jerry Shay (University of Texas Southwestern, Dallas, TX, USA) for providing invaluable guidance on the telomerase and telomere measurement experiments. Davide Pellacani provided many useful discussions concerning the work.
Morbidity and loss of quality of life due to benign prostatic hyperplasia (BPH) are relatively high. There are >210 million cases of BPH worldwide [1], and the histologic prevalence of BPH is approximately 80% at the age of 80 yr. BPH is characterised by uncontrolled but noninvasive growth of a variety of cell types in the prostate, ranging in frequency from adenomyofibromatous, fibromuscular, fibroadenomatous, stromal, to rare muscular compositions. Medical treatments for BPH include targeting of the smooth muscle cells by α1-adrenergic receptor inhibitor and use of steroid 5α-reductase inhibitors to target the androgen-signalling axis. These treatments take several months to show urinary flow rate benefits but perhaps are ineffective in targeting androgen-independent basal cell proliferation. Surgical intervention by transurethral resection relieves urinary flow obstruction.
In normal somatic cells, repeated cell divisions result in an erosion of the repetitive terminal DNA sequences, the telomeres. The ultimate result of extreme telomere shortening is stress signal–mediated apoptosis or senescence. An understanding of telomerase biology is crucial to comprehend the proliferative dynamics of a tissue. Increased telomerase can be detected in approximately 70–90% of human cancers, including prostate cancer [2]. Although BPH tissue also exhibits a proliferative index two to three times higher than normal prostate [3], telomerase remains undetectable or is only sporadically expressed [2]. The inability to detect telomerase in non–stem-like cells, including the differentiated basal and luminal cells (>95% of the lesion), could have been interpreted as absent or sporadic telomerase expression or activity patterns in previous whole-biopsy analyses [2].
We assessed expression of the RNA (TERC) and protein (TERT) components of telomerase enzyme, telomerase activity, and telomere length in uncultured human BPH-derived stem-like cells (SCs: Lin−/CD31−/EpCAM+/CD133+/CD44+/α2β1hi), transit amplifying cells (TAs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1hi), committed basal cells (CBs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1lo), luminal cells (LCs: Lin−/CD31−/EpCAM+/CD44−/CD24+), and stromal cells (Lin−/CD31−/EpCAM−) (Supplementary Fig. 1, Supplement 1) [4] and [5] freshly purified from BPH tissues (Supplementary Table 1). The expression of NANOG was also higher in SCs (Supplementary Fig. 2). Previously, we reported that TERT expression was undetectable in normal prostate epithelial SCs and TAs [6]. In human BPH, however, the SCs and TAs exhibited significant overexpression of both TERC and TERT compared with the more differentiated CBs and LCs and stromal cells (Fig. 1a and 1b). Telomerase activity was also exclusively restricted to the SC/TA compartment, apart from one CB sample (Fig. 1c). All biopsies were specifically taken from the transitional zone with no detectable prostate cancer lesions on histology from patients with serum prostate-specific antigen levels <4 ng/ml to reduce the possibility of contamination with (telomerase-positive) cancer cells (Supplementary Table 1). TERT expression and telomerase activity were undetectable in stromal cells, indicating that telomerase is not essential for stromal hyperproliferation in BPH—another intriguing disparity in earlier data.

Epithelial cell telomere length measurements revealed that telomerase-expressing SCs and TAs had longer telomeres than those of their immediate differentiated progeny, the telomerase-negative CBs (p < 0.01) (Fig. 1d). The relative telomere length of LCs, however, was almost equivalent to that of SCs and TAs and was always longer than telomere lengths of patient-matched CBs (p < 0.01). Because LCs do not exhibit any detectable telomerase activity (Fig. 1c) and are believed to be derived from basal cells, the paradoxical observation of longer LC telomeres compared with CBs suggests that basal and luminal cells may be derived from distinct SC progenitors.
We stained BPH sections to determine basal cell content (using TP63) and luminal cell content (using NKX3.1) (Fig. 2a) and showed that basal and luminal hyperproliferation can occur together and even independently of each other because the basal:luminal ratio was highly variable, ranging from 0.22 to 2.08 (Fig. 2b, Supplementary Fig. 3), providing further evidence that basal and luminal cells may indeed be derived from different progenitors.

In human prostate, we have previously shown that SCs differentiate into CB cells via TA cells [4], unlike mouse cells, in which the situation is less clear [7]. The lineage of LCs, however, has not been determined in humans, although in vitro we can readily derive luminal cells from basal cells under specific culture conditions [5]. We now propose that basal and luminal cell lineages may develop separately in BPH and speculate that the prostate epithelial SCs can asymmetrically differentiate [8] into distinct basal and luminal progenitor cells, as shown in Figure 2c. In many other stem cell systems, the spindle orientation of progenitor progenies can determine their ultimate identity. The two-progenitor model may assist in understanding BPH pathogenesis, in which each progenitor and its progeny can expand independently or simultaneously to give rise to well-documented [9] areas of basal hyperproliferation, luminal hyperproliferation, or proliferation of both compartments, as shown in Figure 2a. If a basal progenitor cell differentiates into CBs, which retain limited proliferative potential in the absence of detectable telomerase activity, this will result in telomere shortening in CBs, for example, in basal cell hyperplasia, as shown in Figure 2a. In contrast, a luminal progenitor cell population expressing telomerase could expand and give rise to terminally differentiated luminal cells, which do not replicate, maintaining a telomere length similar to that of the luminal progenitors. Our data is in agreement with both Bonkhoff et al [10] and McNeal et al [11] with respect to the existence of a proliferative basal compartment in which the potential luminal progenitors derived from basal SCs exhibit characteristics of both basal and luminal phenotypes, analogous to castration-resistant Nkx3.1-expressing cells in mice [12]. The second implication of our model is that local inhibition of telomerase in BPH could be an alternative therapeutic strategy, as it will inhibit the proliferation both luminal and basal epithelial progenitors but perhaps not that of stromal cells. Because it is suggested that these progenitors function as a reservoir of proliferating epithelial cells in BPH, utilisation of novel and specific telomerase inhibitors such as Imetelstat (Geron Corp, Menlo Park, CA, USA) could prove to be a good adjuvant therapy for the management of BPH.
Author contributions: Norman J. Maitland had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Study concept and design: Maitland, Rane.
Acquisition of data: Rane, Frame, Greener.
Analysis and interpretation of data: Maitland, Rane, Frame.
Drafting of the manuscript: Rane.
Critical revision of the manuscript for important intellectual content: Maitland, Rane, Collins, Berney, Frame.
Statistical analysis: Rane, Frame.
Obtaining funding: Maitland.
Administrative, technical, or material support: Mann, Simms, Collins, Berney.
Supervision: Maitland.
Other (specify): None.
Financial disclosures: Norman J. Maitland certifies that all conflicts of interest, including specific financial interests and relationships and affiliations relevant to the subject matter or materials discussed in the manuscript (eg, employment/affiliation, grants or funding, consultancies, honoraria, stock ownership or options, expert testimony, royalties, or patents filed, received, or pending), are the following: None.
Funding/Support and role of the sponsor: The work was funded by Prostate Cancer UK (G2012-37) and PRO-NEST Marie-Curie network (J.K.R.), and Yorkshire Cancer Research (grant Y257PA, F.M.F. and N.J.M.). The human prostate tissue bank was funded by the Orchid Trust (D.B.). The sponsors were involved in the design and conduct of the study and collection and analysis of the data.
Acknowledgements: We would like to thank all of the patients and the urology surgeons L. Coombes, G. Cooksey and J. Hetherington (Castle Hill Hospital, Cottingham, UK). Thank you also to Megan Nash for immunofluorescence staining and counting and to Jerry Shay (University of Texas Southwestern, Dallas, TX, USA) for providing invaluable guidance on the telomerase and telomere measurement experiments. Davide Pellacani provided many useful discussions concerning the work.
Morbidity and loss of quality of life due to benign prostatic hyperplasia (BPH) are relatively high. There are >210 million cases of BPH worldwide [1], and the histologic prevalence of BPH is approximately 80% at the age of 80 yr. BPH is characterised by uncontrolled but noninvasive growth of a variety of cell types in the prostate, ranging in frequency from adenomyofibromatous, fibromuscular, fibroadenomatous, stromal, to rare muscular compositions. Medical treatments for BPH include targeting of the smooth muscle cells by α1-adrenergic receptor inhibitor and use of steroid 5α-reductase inhibitors to target the androgen-signalling axis. These treatments take several months to show urinary flow rate benefits but perhaps are ineffective in targeting androgen-independent basal cell proliferation. Surgical intervention by transurethral resection relieves urinary flow obstruction.
In normal somatic cells, repeated cell divisions result in an erosion of the repetitive terminal DNA sequences, the telomeres. The ultimate result of extreme telomere shortening is stress signal–mediated apoptosis or senescence. An understanding of telomerase biology is crucial to comprehend the proliferative dynamics of a tissue. Increased telomerase can be detected in approximately 70–90% of human cancers, including prostate cancer [2]. Although BPH tissue also exhibits a proliferative index two to three times higher than normal prostate [3], telomerase remains undetectable or is only sporadically expressed [2]. The inability to detect telomerase in non–stem-like cells, including the differentiated basal and luminal cells (>95% of the lesion), could have been interpreted as absent or sporadic telomerase expression or activity patterns in previous whole-biopsy analyses [2].
We assessed expression of the RNA (TERC) and protein (TERT) components of telomerase enzyme, telomerase activity, and telomere length in uncultured human BPH-derived stem-like cells (SCs: Lin−/CD31−/EpCAM+/CD133+/CD44+/α2β1hi), transit amplifying cells (TAs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1hi), committed basal cells (CBs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1lo), luminal cells (LCs: Lin−/CD31−/EpCAM+/CD44−/CD24+), and stromal cells (Lin−/CD31−/EpCAM−) (Supplementary Fig. 1, Supplement 1) [4] and [5] freshly purified from BPH tissues (Supplementary Table 1). The expression of NANOG was also higher in SCs (Supplementary Fig. 2). Previously, we reported that TERT expression was undetectable in normal prostate epithelial SCs and TAs [6]. In human BPH, however, the SCs and TAs exhibited significant overexpression of both TERC and TERT compared with the more differentiated CBs and LCs and stromal cells (Fig. 1a and 1b). Telomerase activity was also exclusively restricted to the SC/TA compartment, apart from one CB sample (Fig. 1c). All biopsies were specifically taken from the transitional zone with no detectable prostate cancer lesions on histology from patients with serum prostate-specific antigen levels <4 ng/ml to reduce the possibility of contamination with (telomerase-positive) cancer cells (Supplementary Table 1). TERT expression and telomerase activity were undetectable in stromal cells, indicating that telomerase is not essential for stromal hyperproliferation in BPH—another intriguing disparity in earlier data.

Epithelial cell telomere length measurements revealed that telomerase-expressing SCs and TAs had longer telomeres than those of their immediate differentiated progeny, the telomerase-negative CBs (p < 0.01) (Fig. 1d). The relative telomere length of LCs, however, was almost equivalent to that of SCs and TAs and was always longer than telomere lengths of patient-matched CBs (p < 0.01). Because LCs do not exhibit any detectable telomerase activity (Fig. 1c) and are believed to be derived from basal cells, the paradoxical observation of longer LC telomeres compared with CBs suggests that basal and luminal cells may be derived from distinct SC progenitors.
We stained BPH sections to determine basal cell content (using TP63) and luminal cell content (using NKX3.1) (Fig. 2a) and showed that basal and luminal hyperproliferation can occur together and even independently of each other because the basal:luminal ratio was highly variable, ranging from 0.22 to 2.08 (Fig. 2b, Supplementary Fig. 3), providing further evidence that basal and luminal cells may indeed be derived from different progenitors.

In human prostate, we have previously shown that SCs differentiate into CB cells via TA cells [4], unlike mouse cells, in which the situation is less clear [7]. The lineage of LCs, however, has not been determined in humans, although in vitro we can readily derive luminal cells from basal cells under specific culture conditions [5]. We now propose that basal and luminal cell lineages may develop separately in BPH and speculate that the prostate epithelial SCs can asymmetrically differentiate [8] into distinct basal and luminal progenitor cells, as shown in Figure 2c. In many other stem cell systems, the spindle orientation of progenitor progenies can determine their ultimate identity. The two-progenitor model may assist in understanding BPH pathogenesis, in which each progenitor and its progeny can expand independently or simultaneously to give rise to well-documented [9] areas of basal hyperproliferation, luminal hyperproliferation, or proliferation of both compartments, as shown in Figure 2a. If a basal progenitor cell differentiates into CBs, which retain limited proliferative potential in the absence of detectable telomerase activity, this will result in telomere shortening in CBs, for example, in basal cell hyperplasia, as shown in Figure 2a. In contrast, a luminal progenitor cell population expressing telomerase could expand and give rise to terminally differentiated luminal cells, which do not replicate, maintaining a telomere length similar to that of the luminal progenitors. Our data is in agreement with both Bonkhoff et al [10] and McNeal et al [11] with respect to the existence of a proliferative basal compartment in which the potential luminal progenitors derived from basal SCs exhibit characteristics of both basal and luminal phenotypes, analogous to castration-resistant Nkx3.1-expressing cells in mice [12]. The second implication of our model is that local inhibition of telomerase in BPH could be an alternative therapeutic strategy, as it will inhibit the proliferation both luminal and basal epithelial progenitors but perhaps not that of stromal cells. Because it is suggested that these progenitors function as a reservoir of proliferating epithelial cells in BPH, utilisation of novel and specific telomerase inhibitors such as Imetelstat (Geron Corp, Menlo Park, CA, USA) could prove to be a good adjuvant therapy for the management of BPH.
Author contributions: Norman J. Maitland had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Study concept and design: Maitland, Rane.
Acquisition of data: Rane, Frame, Greener.
Analysis and interpretation of data: Maitland, Rane, Frame.
Drafting of the manuscript: Rane.
Critical revision of the manuscript for important intellectual content: Maitland, Rane, Collins, Berney, Frame.
Statistical analysis: Rane, Frame.
Obtaining funding: Maitland.
Administrative, technical, or material support: Mann, Simms, Collins, Berney.
Supervision: Maitland.
Other (specify): None.
Financial disclosures: Norman J. Maitland certifies that all conflicts of interest, including specific financial interests and relationships and affiliations relevant to the subject matter or materials discussed in the manuscript (eg, employment/affiliation, grants or funding, consultancies, honoraria, stock ownership or options, expert testimony, royalties, or patents filed, received, or pending), are the following: None.
Funding/Support and role of the sponsor: The work was funded by Prostate Cancer UK (G2012-37) and PRO-NEST Marie-Curie network (J.K.R.), and Yorkshire Cancer Research (grant Y257PA, F.M.F. and N.J.M.). The human prostate tissue bank was funded by the Orchid Trust (D.B.). The sponsors were involved in the design and conduct of the study and collection and analysis of the data.
Acknowledgements: We would like to thank all of the patients and the urology surgeons L. Coombes, G. Cooksey and J. Hetherington (Castle Hill Hospital, Cottingham, UK). Thank you also to Megan Nash for immunofluorescence staining and counting and to Jerry Shay (University of Texas Southwestern, Dallas, TX, USA) for providing invaluable guidance on the telomerase and telomere measurement experiments. Davide Pellacani provided many useful discussions concerning the work.
Morbidity and loss of quality of life due to benign prostatic hyperplasia (BPH) are relatively high. There are >210 million cases of BPH worldwide [1], and the histologic prevalence of BPH is approximately 80% at the age of 80 yr. BPH is characterised by uncontrolled but noninvasive growth of a variety of cell types in the prostate, ranging in frequency from adenomyofibromatous, fibromuscular, fibroadenomatous, stromal, to rare muscular compositions. Medical treatments for BPH include targeting of the smooth muscle cells by α1-adrenergic receptor inhibitor and use of steroid 5α-reductase inhibitors to target the androgen-signalling axis. These treatments take several months to show urinary flow rate benefits but perhaps are ineffective in targeting androgen-independent basal cell proliferation. Surgical intervention by transurethral resection relieves urinary flow obstruction.
In normal somatic cells, repeated cell divisions result in an erosion of the repetitive terminal DNA sequences, the telomeres. The ultimate result of extreme telomere shortening is stress signal–mediated apoptosis or senescence. An understanding of telomerase biology is crucial to comprehend the proliferative dynamics of a tissue. Increased telomerase can be detected in approximately 70–90% of human cancers, including prostate cancer [2]. Although BPH tissue also exhibits a proliferative index two to three times higher than normal prostate [3], telomerase remains undetectable or is only sporadically expressed [2]. The inability to detect telomerase in non–stem-like cells, including the differentiated basal and luminal cells (>95% of the lesion), could have been interpreted as absent or sporadic telomerase expression or activity patterns in previous whole-biopsy analyses [2].
We assessed expression of the RNA (TERC) and protein (TERT) components of telomerase enzyme, telomerase activity, and telomere length in uncultured human BPH-derived stem-like cells (SCs: Lin−/CD31−/EpCAM+/CD133+/CD44+/α2β1hi), transit amplifying cells (TAs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1hi), committed basal cells (CBs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1lo), luminal cells (LCs: Lin−/CD31−/EpCAM+/CD44−/CD24+), and stromal cells (Lin−/CD31−/EpCAM−) (Supplementary Fig. 1, Supplement 1) [4] and [5] freshly purified from BPH tissues (Supplementary Table 1). The expression of NANOG was also higher in SCs (Supplementary Fig. 2). Previously, we reported that TERT expression was undetectable in normal prostate epithelial SCs and TAs [6]. In human BPH, however, the SCs and TAs exhibited significant overexpression of both TERC and TERT compared with the more differentiated CBs and LCs and stromal cells (Fig. 1a and 1b). Telomerase activity was also exclusively restricted to the SC/TA compartment, apart from one CB sample (Fig. 1c). All biopsies were specifically taken from the transitional zone with no detectable prostate cancer lesions on histology from patients with serum prostate-specific antigen levels <4 ng/ml to reduce the possibility of contamination with (telomerase-positive) cancer cells (Supplementary Table 1). TERT expression and telomerase activity were undetectable in stromal cells, indicating that telomerase is not essential for stromal hyperproliferation in BPH—another intriguing disparity in earlier data.

Epithelial cell telomere length measurements revealed that telomerase-expressing SCs and TAs had longer telomeres than those of their immediate differentiated progeny, the telomerase-negative CBs (p < 0.01) (Fig. 1d). The relative telomere length of LCs, however, was almost equivalent to that of SCs and TAs and was always longer than telomere lengths of patient-matched CBs (p < 0.01). Because LCs do not exhibit any detectable telomerase activity (Fig. 1c) and are believed to be derived from basal cells, the paradoxical observation of longer LC telomeres compared with CBs suggests that basal and luminal cells may be derived from distinct SC progenitors.
We stained BPH sections to determine basal cell content (using TP63) and luminal cell content (using NKX3.1) (Fig. 2a) and showed that basal and luminal hyperproliferation can occur together and even independently of each other because the basal:luminal ratio was highly variable, ranging from 0.22 to 2.08 (Fig. 2b, Supplementary Fig. 3), providing further evidence that basal and luminal cells may indeed be derived from different progenitors.

In human prostate, we have previously shown that SCs differentiate into CB cells via TA cells [4], unlike mouse cells, in which the situation is less clear [7]. The lineage of LCs, however, has not been determined in humans, although in vitro we can readily derive luminal cells from basal cells under specific culture conditions [5]. We now propose that basal and luminal cell lineages may develop separately in BPH and speculate that the prostate epithelial SCs can asymmetrically differentiate [8] into distinct basal and luminal progenitor cells, as shown in Figure 2c. In many other stem cell systems, the spindle orientation of progenitor progenies can determine their ultimate identity. The two-progenitor model may assist in understanding BPH pathogenesis, in which each progenitor and its progeny can expand independently or simultaneously to give rise to well-documented [9] areas of basal hyperproliferation, luminal hyperproliferation, or proliferation of both compartments, as shown in Figure 2a. If a basal progenitor cell differentiates into CBs, which retain limited proliferative potential in the absence of detectable telomerase activity, this will result in telomere shortening in CBs, for example, in basal cell hyperplasia, as shown in Figure 2a. In contrast, a luminal progenitor cell population expressing telomerase could expand and give rise to terminally differentiated luminal cells, which do not replicate, maintaining a telomere length similar to that of the luminal progenitors. Our data is in agreement with both Bonkhoff et al [10] and McNeal et al [11] with respect to the existence of a proliferative basal compartment in which the potential luminal progenitors derived from basal SCs exhibit characteristics of both basal and luminal phenotypes, analogous to castration-resistant Nkx3.1-expressing cells in mice [12]. The second implication of our model is that local inhibition of telomerase in BPH could be an alternative therapeutic strategy, as it will inhibit the proliferation both luminal and basal epithelial progenitors but perhaps not that of stromal cells. Because it is suggested that these progenitors function as a reservoir of proliferating epithelial cells in BPH, utilisation of novel and specific telomerase inhibitors such as Imetelstat (Geron Corp, Menlo Park, CA, USA) could prove to be a good adjuvant therapy for the management of BPH.
Author contributions: Norman J. Maitland had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Study concept and design: Maitland, Rane.
Acquisition of data: Rane, Frame, Greener.
Analysis and interpretation of data: Maitland, Rane, Frame.
Drafting of the manuscript: Rane.
Critical revision of the manuscript for important intellectual content: Maitland, Rane, Collins, Berney, Frame.
Statistical analysis: Rane, Frame.
Obtaining funding: Maitland.
Administrative, technical, or material support: Mann, Simms, Collins, Berney.
Supervision: Maitland.
Other (specify): None.
Financial disclosures: Norman J. Maitland certifies that all conflicts of interest, including specific financial interests and relationships and affiliations relevant to the subject matter or materials discussed in the manuscript (eg, employment/affiliation, grants or funding, consultancies, honoraria, stock ownership or options, expert testimony, royalties, or patents filed, received, or pending), are the following: None.
Funding/Support and role of the sponsor: The work was funded by Prostate Cancer UK (G2012-37) and PRO-NEST Marie-Curie network (J.K.R.), and Yorkshire Cancer Research (grant Y257PA, F.M.F. and N.J.M.). The human prostate tissue bank was funded by the Orchid Trust (D.B.). The sponsors were involved in the design and conduct of the study and collection and analysis of the data.
Acknowledgements: We would like to thank all of the patients and the urology surgeons L. Coombes, G. Cooksey and J. Hetherington (Castle Hill Hospital, Cottingham, UK). Thank you also to Megan Nash for immunofluorescence staining and counting and to Jerry Shay (University of Texas Southwestern, Dallas, TX, USA) for providing invaluable guidance on the telomerase and telomere measurement experiments. Davide Pellacani provided many useful discussions concerning the work.
Morbidity and loss of quality of life due to benign prostatic hyperplasia (BPH) are relatively high. There are >210 million cases of BPH worldwide [1], and the histologic prevalence of BPH is approximately 80% at the age of 80 yr. BPH is characterised by uncontrolled but noninvasive growth of a variety of cell types in the prostate, ranging in frequency from adenomyofibromatous, fibromuscular, fibroadenomatous, stromal, to rare muscular compositions. Medical treatments for BPH include targeting of the smooth muscle cells by α1-adrenergic receptor inhibitor and use of steroid 5α-reductase inhibitors to target the androgen-signalling axis. These treatments take several months to show urinary flow rate benefits but perhaps are ineffective in targeting androgen-independent basal cell proliferation. Surgical intervention by transurethral resection relieves urinary flow obstruction.
In normal somatic cells, repeated cell divisions result in an erosion of the repetitive terminal DNA sequences, the telomeres. The ultimate result of extreme telomere shortening is stress signal–mediated apoptosis or senescence. An understanding of telomerase biology is crucial to comprehend the proliferative dynamics of a tissue. Increased telomerase can be detected in approximately 70–90% of human cancers, including prostate cancer [2]. Although BPH tissue also exhibits a proliferative index two to three times higher than normal prostate [3], telomerase remains undetectable or is only sporadically expressed [2]. The inability to detect telomerase in non–stem-like cells, including the differentiated basal and luminal cells (>95% of the lesion), could have been interpreted as absent or sporadic telomerase expression or activity patterns in previous whole-biopsy analyses [2].
We assessed expression of the RNA (TERC) and protein (TERT) components of telomerase enzyme, telomerase activity, and telomere length in uncultured human BPH-derived stem-like cells (SCs: Lin−/CD31−/EpCAM+/CD133+/CD44+/α2β1hi), transit amplifying cells (TAs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1hi), committed basal cells (CBs: Lin−/CD31−/EpCAM+/CD133−/CD44+/α2β1lo), luminal cells (LCs: Lin−/CD31−/EpCAM+/CD44−/CD24+), and stromal cells (Lin−/CD31−/EpCAM−) (Supplementary Fig. 1, Supplement 1) [4] and [5] freshly purified from BPH tissues (Supplementary Table 1). The expression of NANOG was also higher in SCs (Supplementary Fig. 2). Previously, we reported that TERT expression was undetectable in normal prostate epithelial SCs and TAs [6]. In human BPH, however, the SCs and TAs exhibited significant overexpression of both TERC and TERT compared with the more differentiated CBs and LCs and stromal cells (Fig. 1a and 1b). Telomerase activity was also exclusively restricted to the SC/TA compartment, apart from one CB sample (Fig. 1c). All biopsies were specifically taken from the transitional zone with no detectable prostate cancer lesions on histology from patients with serum prostate-specific antigen levels <4 ng/ml to reduce the possibility of contamination with (telomerase-positive) cancer cells (Supplementary Table 1). TERT expression and telomerase activity were undetectable in stromal cells, indicating that telomerase is not essential for stromal hyperproliferation in BPH—another intriguing disparity in earlier data.

Epithelial cell telomere length measurements revealed that telomerase-expressing SCs and TAs had longer telomeres than those of their immediate differentiated progeny, the telomerase-negative CBs (p < 0.01) (Fig. 1d). The relative telomere length of LCs, however, was almost equivalent to that of SCs and TAs and was always longer than telomere lengths of patient-matched CBs (p < 0.01). Because LCs do not exhibit any detectable telomerase activity (Fig. 1c) and are believed to be derived from basal cells, the paradoxical observation of longer LC telomeres compared with CBs suggests that basal and luminal cells may be derived from distinct SC progenitors.
We stained BPH sections to determine basal cell content (using TP63) and luminal cell content (using NKX3.1) (Fig. 2a) and showed that basal and luminal hyperproliferation can occur together and even independently of each other because the basal:luminal ratio was highly variable, ranging from 0.22 to 2.08 (Fig. 2b, Supplementary Fig. 3), providing further evidence that basal and luminal cells may indeed be derived from different progenitors.

In human prostate, we have previously shown that SCs differentiate into CB cells via TA cells [4], unlike mouse cells, in which the situation is less clear [7]. The lineage of LCs, however, has not been determined in humans, although in vitro we can readily derive luminal cells from basal cells under specific culture conditions [5]. We now propose that basal and luminal cell lineages may develop separately in BPH and speculate that the prostate epithelial SCs can asymmetrically differentiate [8] into distinct basal and luminal progenitor cells, as shown in Figure 2c. In many other stem cell systems, the spindle orientation of progenitor progenies can determine their ultimate identity. The two-progenitor model may assist in understanding BPH pathogenesis, in which each progenitor and its progeny can expand independently or simultaneously to give rise to well-documented [9] areas of basal hyperproliferation, luminal hyperproliferation, or proliferation of both compartments, as shown in Figure 2a. If a basal progenitor cell differentiates into CBs, which retain limited proliferative potential in the absence of detectable telomerase activity, this will result in telomere shortening in CBs, for example, in basal cell hyperplasia, as shown in Figure 2a. In contrast, a luminal progenitor cell population expressing telomerase could expand and give rise to terminally differentiated luminal cells, which do not replicate, maintaining a telomere length similar to that of the luminal progenitors. Our data is in agreement with both Bonkhoff et al [10] and McNeal et al [11] with respect to the existence of a proliferative basal compartment in which the potential luminal progenitors derived from basal SCs exhibit characteristics of both basal and luminal phenotypes, analogous to castration-resistant Nkx3.1-expressing cells in mice [12]. The second implication of our model is that local inhibition of telomerase in BPH could be an alternative therapeutic strategy, as it will inhibit the proliferation both luminal and basal epithelial progenitors but perhaps not that of stromal cells. Because it is suggested that these progenitors function as a reservoir of proliferating epithelial cells in BPH, utilisation of novel and specific telomerase inhibitors such as Imetelstat (Geron Corp, Menlo Park, CA, USA) could prove to be a good adjuvant therapy for the management of BPH.
Author contributions: Norman J. Maitland had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Study concept and design: Maitland, Rane.
Acquisition of data: Rane, Frame, Greener.
Analysis and interpretation of data: Maitland, Rane, Frame.
Drafting of the manuscript: Rane.
Critical revision of the manuscript for important intellectual content: Maitland, Rane, Collins, Berney, Frame.
Statistical analysis: Rane, Frame.
Obtaining funding: Maitland.
Administrative, technical, or material support: Mann, Simms, Collins, Berney.
Supervision: Maitland.
Other (specify): None.
Financial disclosures: Norman J. Maitland certifies that all conflicts of interest, including specific financial interests and relationships and affiliations relevant to the subject matter or materials discussed in the manuscript (eg, employment/affiliation, grants or funding, consultancies, honoraria, stock ownership or options, expert testimony, royalties, or patents filed, received, or pending), are the following: None.
Funding/Support and role of the sponsor: The work was funded by Prostate Cancer UK (G2012-37) and PRO-NEST Marie-Curie network (J.K.R.), and Yorkshire Cancer Research (grant Y257PA, F.M.F. and N.J.M.). The human prostate tissue bank was funded by the Orchid Trust (D.B.). The sponsors were involved in the design and conduct of the study and collection and analysis of the data.
Acknowledgements: We would like to thank all of the patients and the urology surgeons L. Coombes, G. Cooksey and J. Hetherington (Castle Hill Hospital, Cottingham, UK). Thank you also to Megan Nash for immunofluorescence staining and counting and to Jerry Shay (University of Texas Southwestern, Dallas, TX, USA) for providing invaluable guidance on the telomerase and telomere measurement experiments. Davide Pellacani provided many useful discussions concerning the work.